使用我的GSE157827数据进行初步分析-3
other-other2026.1.28-2026.3.11研究进展
hERV计数
stellarscope:使用best_average模式
module load miniconda3/base
conda activate stellarscope
bam_dir="/public/home/GENE_proc/wth/GSE157827/bam/filter_bam"
stellarscope_dir="/public/home/GENE_proc/wth/GSE157827/stellarscope"
out_root="/public/home/GENE_proc/wth/GSE157827/mtx"
hERV_gtf=/public/home/wangtianhao/Desktop/STAR_ref/transcripts.gtf
for bam in "${bam_dir}"/GSM*Aligned.sortedByCB.bam; do
GSM=$(basename "$bam" | sed 's/Aligned\.sortedByCB\.bam$//')
outdir="${stellarscope_dir}/${GSM}/"
mkdir -p "$outdir"
stellarscope assign \
--outdir "$outdir" \
--nproc 16 \
--stranded_mode F \
--whitelist "${out_root}/${GSM}/gene/barcodes.tsv" \
--pooling_mode individual \
--reassign_mode best_average \
--max_iter 500 \
--updated_sam \
"$bam" \
"$hERV_gtf"
mkdir -p ${out_root}/${GSM}/hERV_new
cp ${outdir}/stellarscope-barcodes.tsv ${out_root}/${GSM}/hERV_new/barcodes.tsv
cp ${outdir}/stellarscope-features.tsv ${out_root}/${GSM}/hERV_new/features.tsv
cp ${outdir}/stellarscope-TE_counts.mtx ${out_root}/${GSM}/hERV_new/counts.mtx
done
补充:计数结果feature中的__no_feature是什么?
- 不是一个真实的TE位点/家族,而是一个“未能匹配到任何注释feature的类别”
- 通常后续分析可以忽略这项
featureCounts:不提供直接适配单细胞数据的接口(不会按barcode分列,也不做UMI去重),需要结合umi-tools等工具处理(参见单细胞转录组数据分析流程,但这个教程也是针对普通基因的)
scTE:是适配单细胞数据的转座元件分析工具,不过也是需要自己做针对hERV的gtf,相比featureCounts可行性更高
得到的mtx文件比原来的大10%~20%。如果原来的比较小(100K左右),新文件可能比原来的大50%
计算family-level计数矩阵
构建映射
构建一个位点-家族-亚家族的映射
get_attr <- function(x, key) {
pat <- paste0(key, ' "([^"]+)"')
m <- regexec(pat, x, perl = TRUE)
rr <- regmatches(x, m)
out <- rep(NA_character_, length(x))
ok <- lengths(rr) >= 2
out[ok] <- vapply(rr[ok], `[`, character(1), 2)
out
}
build_herv_map <- function(gtf_path) {
lines <- readLines(gtf_path, warn = FALSE)
lines <- lines[!grepl("^\\s*#", lines)]
lines <- lines[nzchar(lines)]
gtf <- data.table::fread(
text = lines,
sep = "\t",
header = FALSE,
quote = "",
fill = TRUE,
data.table = TRUE
)
gtf <- gtf[, 1:9]
setnames(gtf, c("seqname","source","feature","start","end","score","strand","frame","attribute"))
gtf[, gene_id := get_attr(attribute, "gene_id")]
gtf[, locus := get_attr(attribute, "locus")]
gtf[, intModel := get_attr(attribute, "intModel")]
gtf[, repName := get_attr(attribute, "repName")]
gtf[, repFamily := get_attr(attribute, "repFamily")]
gtf[, geneRegion := get_attr(attribute, "geneRegion")]
# gene行:intModel
gene_dt <- gtf[feature == "gene", .(locus_id = gene_id, intModel)][!is.na(locus_id)]
# exon行:repFamily/repName/geneRegion
exon_dt <- gtf[feature == "exon", .(locus_id = gene_id, repFamily, repName, geneRegion)][!is.na(locus_id)]
# 每个位点的repFamily:取出现最多的repFamily
repfam_dt <- exon_dt[!is.na(repFamily),
.(repFamily = {
x <- repFamily
ux <- unique(x); ux[which.max(tabulate(match(x, ux)))]
}),
by = locus_id]
# subfamily:优先internal exon的repName
subfam_dt <- exon_dt[geneRegion == "internal" & !is.na(repName),
.(subfamily_fallback = repName[1]),
by = locus_id]
map <- merge(gene_dt, repfam_dt, by = "locus_id", all = TRUE)
map <- merge(map, subfam_dt, by = "locus_id", all = TRUE)
map[, family := sub("_.*$", "", locus_id)]
map[, subfamily := fifelse(!is.na(intModel) & nzchar(intModel), intModel, subfamily_fallback)]
map$subfamily <- gsub("-int", "", map$subfamily) # 去掉结尾的-int标记
unique(map[, .(locus_id, family, subfamily, repFamily)])
}
gtf_path <- "C:\\Users\\17185\\Desktop\\hERV_calc\\GSE157827\\metadata\\hERV.gtf"
herv_map <- build_herv_map(gtf_path)
# subfamily->family?
normalize_subfamily <- function(x) {
x %>%
str_replace("-int$", "") %>%
str_replace("_I$", "") %>%
str_replace("_a$", "")
}
subfamily_to_family_new <- function(subfamily) {
x <- normalize_subfamily(subfamily)
x <- recode(
x,
"HERVIP10FH" = "HERVIP10F",
.default = x
)
x <- if_else(str_detect(x, "^ERVL"), "ERVL", x)
x <- if_else(str_detect(x, "^HERVK"), "HERVK", x)
x <- if_else(str_detect(x, "^HERVL"), "HERVL", x)
x <- if_else(str_detect(x, "^HERVH"), "HERVH", x)
x <- if_else(str_detect(x, "^HERV-Fc"), "HERV-Fc", x)
x <- if_else(str_detect(x, "^HERVFH"), "HERVFH", x)
x <- if_else(str_detect(x, "^HUERS"), "HUERS", x)
x <- if_else(str_detect(x, "^PRIMA"), "PRIMA", x)
x <- if_else(str_detect(x, "^PABL"), "PABL", x)
x
}
df <- herv_map %>%
mutate(family_new = subfamily_to_family_new(subfamily))
write.csv(df, file = "C:\\Users\\17185\\Desktop\\hERV_calc\\GSE157827\\metadata\\hERV_locus2family.csv", row.names = F)
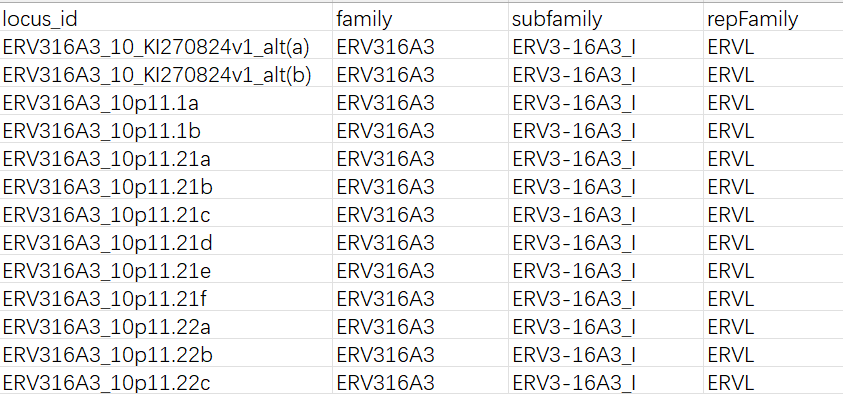

gtf文件:
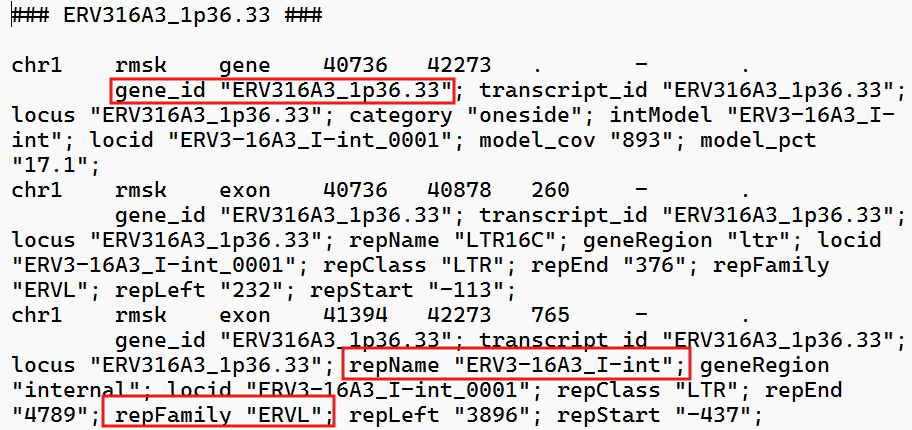
各列含义:
locus_id:计数直接得到的feature名,对应gtf中的gene_idrepFamily:gtf中的repFamily,应该是超家族/大类,只有ERVK/ERVL/ERV1这三种-
subfamily:gtf中的intModel/repName,其中以gene行的intModel优先(如果这个位点在GTF的gene记录里有intModel,就认为它是该位点最核心、最想表达的“内部模型/亚家族标签”,从上面的截图中可以看到,有时一个位点还会有LTR的标记,不过由于计数结果中并没有标明某个读段到底是ltr还是int,就统一都拿int的名字来指代了。如果没有int就用LTR名称在处理时,因为它们都以
-int结尾(表示internal模块),为了更美观将其去掉,只保留前面的家族名 -
family:使用上次的方法,直接从feature名中以-为分隔截取的经过对比,其实subfamily和前面直接按
-分隔的结果差不多,主要的区别是有些subfamily的名称不一致,比如HML系列:- HML1-HERVK14
- HML2-HERVK
- HML3-HERVK9
- HML4-HERVK13
- HML5-HERVK22
- HML6-HERVK3
最终得到的family/subfamily名:
ERV3-16A3_I ERVL-B4 ERVL-E ERVL
Harlequin HERV30 HERV3 HERV4_I
HERV9 HERVE_a HERVE HERV-Fc1
HERV-Fc2 HERVFH19 HERVFH21 MER50
HERVH48 HERVH HERVIP10FH HERVIP10F
HERVI HERVK11D HERVK11 HERVK14C
HERVKC4 HERVL18 HERVL32 HERVL40
HERVL66 HERVL74 HERVL HERVP71A
HERVS71 HERV17 HERVK14 HERVK
HERVK9 HERVK13 HERVK22 HERVK3
HUERS-P1 HUERS-P2 HUERS-P3b HUERS-P3
LTR19 LTR23 LTR25 LTR46
LTR57 MER101 MER34B MER41
MER4B MER4 MER61 PABL_A
PABL_B PRIMA41 PRIMA4 PRIMAX
我还找了当时用featureCounts得到的结果,其实家族名也都是
HERV16 HERV9N HERVIP10F HERVL HERVK13
HERVL40 HERVH HERVL66 HERVFH21 HERV9
HERVFH19 HERVIP10FH HERVK HERV3 HERVE
HERV9NC HERVL74 HERV HERV35I HERVK14C
HERV1 HERV4 HERVK22 HERV30 HERVS71
HERVK11 HERV17 HERVIP10B3 HERVK14 HERVL18
HERVP71A HERVL32 HERVH48 HERVKC4 HERVI
HERVK3 HERVK11D HERV15 HERVK9
这种,没有特别标准的HERVK/HERVH/HERVL……这种聚合(featureCounts用的是repeatmasker官网下载的gtf,我自己提取了名字以HERV开头的行,这里我用的stellarscope作者自己整理的gtf,会包含一些ltr以及一些看起来不像HERV的条目)
一个问题:subfamily看起来很杂乱,repfamily又只有这三类,有没有一个中间层级?
- 在ucsc中搜索subfamily:PRIMA4-int的示例结果,其中的family似乎就是ERV1
- 在dfam中搜索subfamily:HERVE的示例结果,注意description那一栏,ERV1后直接就是不同的subfamily了。而结果中
PABL_B-int这种名字不包括HERVE的,能被搜索出来是因为 点进去之后的summary中提到与HERVE相关(“This is related to the HERV3/HERV4/HERVE group of ERVs”/”it was probably dependent on HERVE_a”),但也没有提到归属关系 - 而gtf文件中更没有相关记述,所以就难以找到中间层级,因此我只能试着去掉后缀(
-a/-A这些),并按前缀聚合,不过因为这个是没有依据的,所以后面我也不太想用这个层级
ERV3-16A3 ERVL Harlequin HERV30 HERV3
HERV4 HERV9 HERVE HERV-Fc HERVFH
MER50 HERVH HERVIP10F HERVI HERVK
HERVL HERVP71A HERVS71 HERV17 HUERS
LTR19 LTR23 LTR25 LTR46 LTR57
MER101 MER34B MER41 MER4B MER4
MER61 PABL PRIMA
替换原assay
注:因为前面构建映射时没有把--no-feature这个feature写进去,所以后面聚合时,会自动把这个过滤掉,而hERV位点的assay是直接用原始计数矩阵导入的,因此nCount_HERV和nCount_HERV_family不同
# meta信息
metadata_xlsx <- "/public/home/GENE_proc/wth/GSE157827/raw_data/metadata/metadata.xlsx"
sample_info_xlsx <- "/public/home/GENE_proc/wth/GSE157827/raw_data/metadata/sample_info.xlsx"
meta_map <- readxl::read_xlsx(metadata_xlsx)
sample_info <- readxl::read_xlsx(sample_info_xlsx)
meta_map <- meta_map |> select(individuals, srr_id, sample_id, tissue)
colnames(meta_map) <- c("sample_id","SRR_id","GSM","tissue")
metadata <- meta_map |> left_join(sample_info, by="sample_id")
gsm2sample <- metadata |> distinct(GSM, sample_id)
# 读取计数矩阵
data_root <- "/public/home/GENE_proc/wth/GSE157827/mtx"
seu <- readRDS("/public/home/GENE_proc/wth/GSE157827/my_data/GSE157827_celltype.rds")
normalize_barcode <- function(x) {
ifelse(grepl("-\\d+$", x), x, paste0(x, "-1"))
}
read_one_gsm_herv_new <- function(GSM){
herv_dir <- file.path(data_root, GSM, "hERV_new")
mat <- ReadMtx(
mtx = file.path(herv_dir, "counts.mtx"),
features = file.path(herv_dir, "features.tsv"),
cells = file.path(herv_dir, "barcodes.tsv"),
feature.column = 1
)
sample_name <- gsm2sample$sample_id[gsm2sample$GSM == GSM][1]
colnames(mat) <- paste(GSM, sample_name, normalize_barcode(colnames(mat)), sep = "_")
mat
}
gsm_ids <- unique(seu$GSM)
herv_list <- lapply(gsm_ids, read_one_gsm_herv_new)
herv_counts_new <- do.call(cbind, herv_list)
# 家族级聚合
herv_map <- read.csv("/public/home/GENE_proc/wth/GSE157827/raw_data/metadata/hERV_locus2family.csv")
aggregate_herv_counts <- function(herv_counts, map, group_by = c("family","subfamily","repFamily", "family_new")) {
group_by <- match.arg(group_by)
keep_loci <- intersect(rownames(herv_counts), map$locus_id)
map2 <- map[match(keep_loci, map$locus_id), ]
map2 <- map2[!is.na(map2[[group_by]]) & nzchar(map2[[group_by]]), ]
mat <- herv_counts[map2$locus_id, , drop = FALSE]
grp <- map2[[group_by]]
f <- factor(grp, levels = unique(grp))
G <- sparseMatrix(
i = seq_along(f),
j = as.integer(f),
x = 1,
dims = c(length(f), nlevels(f)),
dimnames = list(rownames(mat), levels(f))
)
out <- Matrix::t(G) %*% mat
out
}
herv_family_counts_new <- aggregate_herv_counts(herv_counts_new, herv_map, group_by = "subfamily")
herv_repfamily_counts_new <- aggregate_herv_counts(herv_counts_new, herv_map, group_by = "repFamily")
herv_family_new_counts_new <- aggregate_herv_counts(herv_counts_new, herv_map, group_by = "family_new")
# 将新计数矩阵的细胞与原来的对齐
align_to_cells <- function(mat, cells){
mat <- mat[, cells, drop = FALSE]
mat
}
cells <- colnames(seu)
herv_counts_new <- align_to_cells(herv_counts_new, cells)
herv_family_counts_new <- align_to_cells(herv_family_counts_new, cells)
herv_repfamily_counts_new <- align_to_cells(herv_repfamily_counts_new, cells)
herv_family_new_counts_new <- align_to_cells(herv_family_new_counts_new, cells)
# 替换原assay
replace_assay <- function(seu, assay_name, counts){
counts <- as(counts, "dgCMatrix")
seu[[assay_name]] <- CreateAssayObject(counts = counts)
seu
}
seu <- replace_assay(seu, "HERV", herv_counts_new)
seu <- replace_assay(seu, "HERV_family", herv_family_counts_new)
seu <- replace_assay(seu, "HERV_repfamily", herv_repfamily_counts_new)
seu <- replace_assay(seu, "HERV_family_new", herv_family_new_counts_new)
# 更新metadata
herv_counts <- GetAssayData(seu, assay="HERV", slot="counts")
rna_counts <- GetAssayData(seu, assay="RNA", slot="counts")
seu$nCount_HERV <- Matrix::colSums(herv_counts)
seu$nFeature_HERV <- Matrix::colSums(herv_counts > 0)
seu$nCount_HERV_family <- Matrix::colSums(GetAssayData(seu, assay="HERV_family", slot="counts"))
seu$nFeature_HERV_family <- Matrix::colSums(GetAssayData(seu, assay="HERV_family", slot="counts") > 0)
seu$HERV_fraction <- seu$nCount_HERV / Matrix::colSums(rna_counts)
# 标准化
normalize_herv_by_rna_depth <- function(seu, assay, scale.factor = 10000) {
rna_counts <- GetAssayData(seu, assay = "RNA", slot = "counts")
herv_counts <- GetAssayData(seu, assay = assay, slot = "counts")
lib_rna <- Matrix::colSums(rna_counts)
herv_norm <- t(t(herv_counts) / lib_rna) * scale.factor
herv_norm@x <- log1p(herv_norm@x)
seu <- SetAssayData(seu, assay = assay, slot = "data", new.data = herv_norm)
return(seu)
}
seu <- normalize_herv_by_rna_depth(seu, "HERV")
seu <- normalize_herv_by_rna_depth(seu, "HERV_family")
seu <- normalize_herv_by_rna_depth(seu, "HERV_repfamily")
seu <- normalize_herv_by_rna_depth(seu, "HERV_family_new")
# 保存数据
DefaultAssay(seu) <- "RNA"
saveRDS(
seu,
file = "/public/home/GENE_proc/wth/GSE157827/my_data/GSE157827_celltype_new.rds",
compress = "xz"
)
共包含5个assay:
RNA:普通基因(STAR的输出)HERV:hERV位点级计数(stellarscope的输出)HERV_family:subfamily级计数(根据gtf文件)HERV_family_new:family级计数(根据前缀合并到更大的层级)HERV_repfamily:超家族/class级计数(根据gtf文件)
step1:总体/超家族重调情况
总体思路:
- 把所有hERV位点的计数加在一起
- 从原始count开始累加
- 用RNA库深度标准化
- 把累加后的结果作为一个新feature
HERV-total保存在HERV_repfamily里
- AD-NC的差异分析:沿用之前的思路——
FindMarkers+test.use="MAST"+latent.vars=c("orig.ident","nCount_RNA","percent_mito")- 所有细胞
- 取出各细胞类型的细胞单独做
因为在之前的分析中,我看到hERV-low(偏向AD的细胞)普遍测序深度较低,先看看AD/NC的测序深度有没有显著差别
meta <- seu@meta.data %>%
filter(!is.na(group), !is.na(nCount_RNA), !is.na(celltype)) %>%
mutate(group = factor(group, levels = c("NC","AD")))
# 全体细胞
p1 <- ggplot(meta, aes(x = group, y = nCount_RNA)) +
geom_violin(scale = "width", trim = TRUE) +
stat_compare_means(method = "t.test", label = "p.signif") +
scale_y_log10()
# 各大细胞类型
p2 <- ggplot(meta, aes(x = celltype, y = nCount_RNA, fill = group)) +
geom_violin(scale = "width", trim = TRUE, position = position_dodge(width = 0.9)) +
stat_compare_means(method = "t.test", label = "p.signif") +
scale_y_log10()
p1 | p2
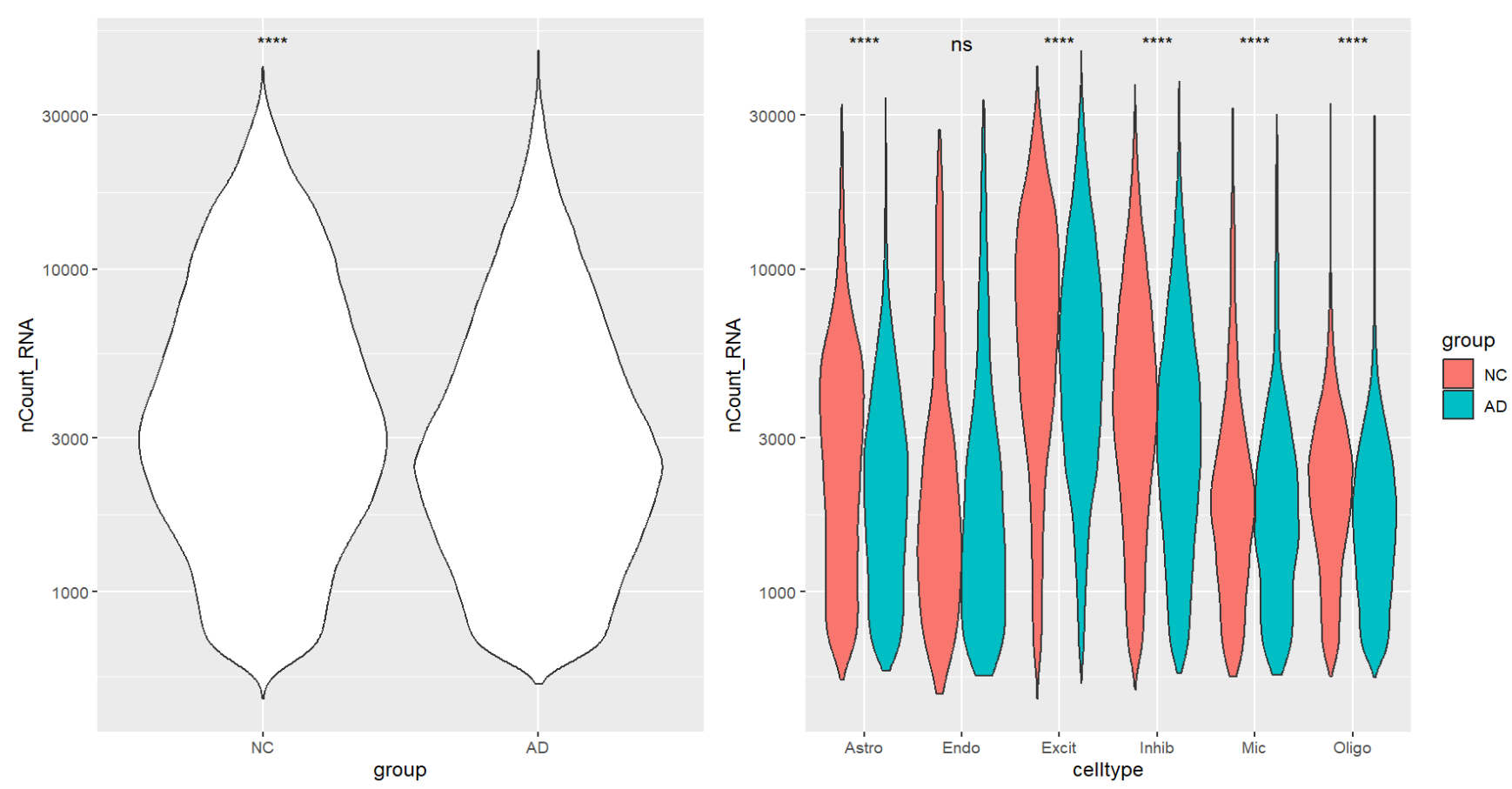
看起来AD的nCount_RNA平均更低,也许前面得到的“AD中hERV表达量更低”的结论是受了测序深度的影响,所以后面尽力用协变量消除一下测序深度的影响吧
- 不过测序深度低是会导致hERV标准化后的值变高还是变低,这个还不太确定,因为我们标准化的方法是
HERV_count / RNA_lib,是count变少的多,还是RNA_lib变少的多
获得总hERV计数:
# hERV计数矩阵
herv_counts <- GetAssayData(seu, assay = "HERV_repfamily", slot = "counts")
herv_total_counts <- Matrix::colSums(herv_counts)
# RNA库深度
rna_counts <- GetAssayData(seu, assay = "RNA", slot = "counts")
rna_lib <- Matrix::colSums(rna_counts)
# 标准化
sf <- 1e4
herv_total_data <- log1p((herv_total_counts / rna_lib) * sf)
# 写入一个新assay
new_counts_row <- Matrix::Matrix(herv_total_counts, nrow = 1, sparse = TRUE)
rownames(new_counts_row) <- "HERV_total"
colnames(new_counts_row) <- colnames(seu)
new_data_row <- Matrix::Matrix(herv_total_data, nrow = 1, sparse = TRUE)
rownames(new_data_row) <- "HERV_total"
colnames(new_data_row) <- colnames(seu)
# 添加到HERV_repfamily的assay中
mat_counts <- GetAssayData(seu, assay = "HERV_repfamily", slot = "counts")
mat_counts2 <- rbind(mat_counts, new_counts_row)
mat_data <- GetAssayData(seu, assay = "HERV_repfamily", slot = "data")
mat_data2 <- rbind(mat_data, new_data_row)
new_assay <- CreateAssayObject(counts = mat_counts2)
new_assay <- SetAssayData(new_assay, slot="data", new.data = mat_data2)
seu[["HERV_repfamily"]] <- new_assay
# 准备差异分析
Idents(seu) <- factor(seu$group, levels = c("NC","AD"))
DefaultAssay(seu) <- "HERV_repfamily"
latent <- c("orig.ident", "nCount_RNA", "percent_mito")
全体细胞:
res_all <- FindMarkers(
object = seu,
ident.1 = "AD", ident.2 = "NC",
test.use = "MAST",
latent.vars = latent,
logfc.threshold = 0, min.pct = 0, return.thresh = 1
)
VlnPlot(seu, features = Features(seu), group.by = "group", pt.size = 0, ncol = 2)
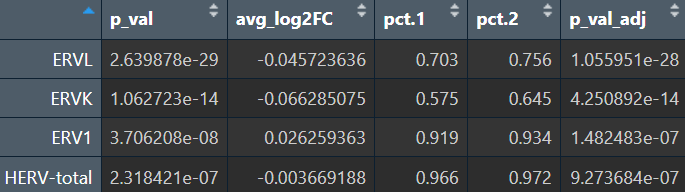
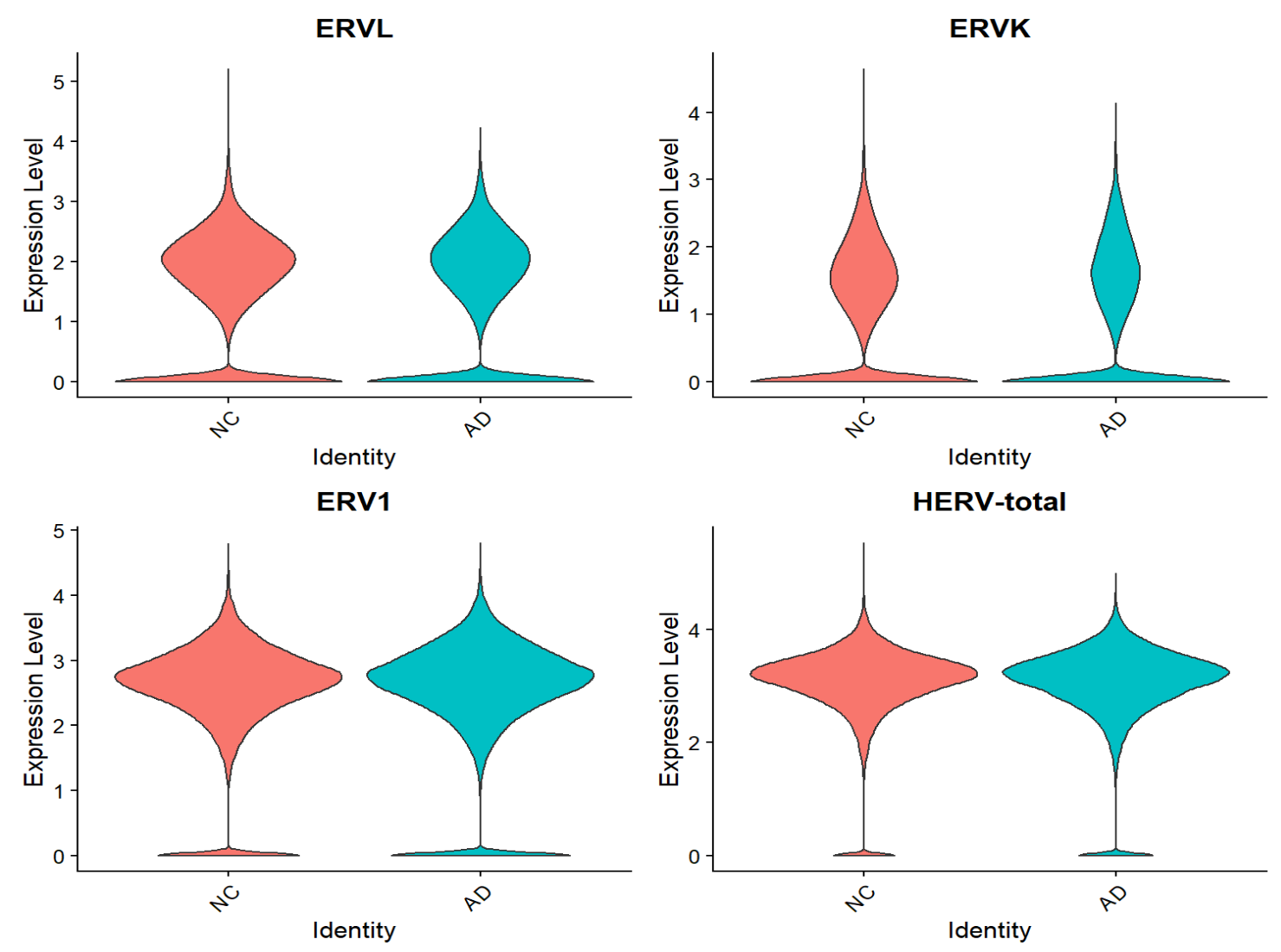
各细胞类型:
celltypes <- sort(unique(seu$celltype))
run_one_ct <- function(ct) {
obj <- subset(seu, subset = celltype == ct)
n_tab <- table(obj$group)
print(paste("running:", ct))
fm <- FindMarkers(
object = obj,
ident.1 = "AD", ident.2 = "NC",
test.use = "MAST",
latent.vars = latent,
logfc.threshold = 0, min.pct = 0, return.thresh = 1
)
out <- fm %>%
tibble::rownames_to_column("feature") %>%
mutate(celltype = ct,
n_NC = unname(n_tab["NC"]),
n_AD = unname(n_tab["AD"]))
out
}
res_by_ct <- do.call(rbind, lapply(celltypes, run_one_ct))
save(res_by_ct, res_all, file = "C:\\Users\\17185\\Desktop\\hERV_calc\\GSE157827\\my_data\\0227_1.RData")
# 只看p值<0.05的
View(res_by_ct %>%
dplyr::filter(p_val_adj<0.05) %>%
arrange(p_val_adj) %>%
select(celltype, n_NC, n_AD, avg_log2FC, p_val, p_val_adj, everything()))
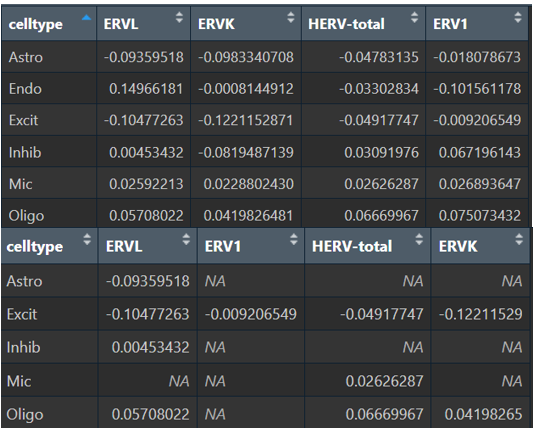
# 全部数据的logFC
View(
res_by_ct %>%
select(celltype, avg_log2FC, feature) %>%
tidyr::pivot_wider(id_cols = celltype, names_from = feature, values_from = avg_log2FC)
)
# 过滤p值后的logFC
View(
res_by_ct %>%
dplyr::filter(p_val_adj<0.05) %>%
select(celltype, avg_log2FC, feature) %>%
tidyr::pivot_wider(id_cols = celltype, names_from = feature, values_from = avg_log2FC,)
)
VlnPlot(seu, features = Features(seu), group.by = "celltype", split.by = "group", pt.size = 0, ncol = 2) +
RotatedAxis()
# 加上图例和显著性标记
mat4 <- GetAssayData(seu, assay = "HERV_repfamily", slot = "data")[c("ERVL","ERVK","ERV1" ,"HERV-total"), , drop = FALSE]
meta <- seu@meta.data %>%
transmute(
cell = rownames(seu@meta.data),
group = factor(group, levels = c("NC","AD")),
celltype = factor(celltype, levels = c("Astro","Endo","Excit","Inhib","Mic","Oligo"))
)
df_expr <- as.data.frame(as.matrix(t(mat4))) %>%
tibble::rownames_to_column("cell") %>%
left_join(meta, by = "cell") %>%
pivot_longer(cols = c("ERVL","ERVK","ERV1","HERV-total"),
names_to = "feature", values_to = "expr") %>%
mutate(
feature = recode(feature, HERV_total = "HERV-total")
)
p_to_star <- function(p){
dplyr::case_when(
is.na(p) ~ "",
p < 0.001 ~ "***",
p < 0.01 ~ "**",
p < 0.05 ~ "*",
TRUE ~ ""
)
}
res_stars <- res_by_ct %>%
mutate(feature = recode(feature, HERV_total = "HERV-total")) %>%
select(celltype, feature, p_val_adj) %>%
distinct() %>%
mutate(star = p_to_star(p_val_adj)) %>%
filter(star != "")
ymax_df <- df_expr %>%
group_by(feature, celltype) %>%
summarise(ymax = max(expr, na.rm = TRUE), .groups = "drop")
anno_df <- res_stars %>%
inner_join(ymax_df, by = c("feature","celltype")) %>%
mutate(y = ymax + 0.15)
cols <- c("NC" = "#F8766D", "AD" = "#00BFC4")
ggplot(df_expr, aes(x = celltype, y = expr, fill = group)) +
geom_violin(
position = position_dodge(width = 0.85),
trim = TRUE, scale = "width", linewidth = 0.25
) +
facet_wrap(~ feature, ncol = 2, scales = "free_y") +
scale_fill_manual(values = cols, name = "Group") +
labs(x = "Cell type", y = "Expression level") +
theme_bw(base_size = 12) +
theme(
legend.position = "right",
axis.text.x = element_text(angle = 45, hjust = 1),
panel.grid = element_blank(),
strip.text = element_text(face = "bold")
) +
geom_text(
data = anno_df,
aes(x = celltype, y = y, label = star),
inherit.aes = FALSE,
size = 5,
vjust = 0
)
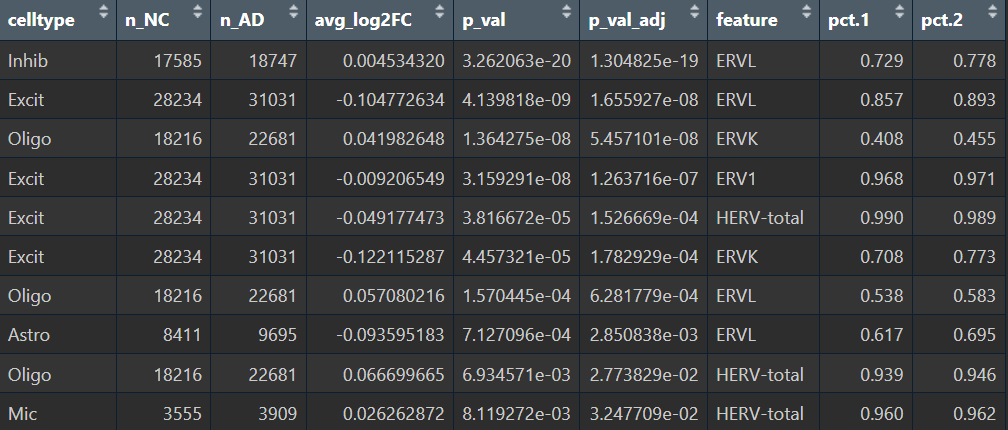
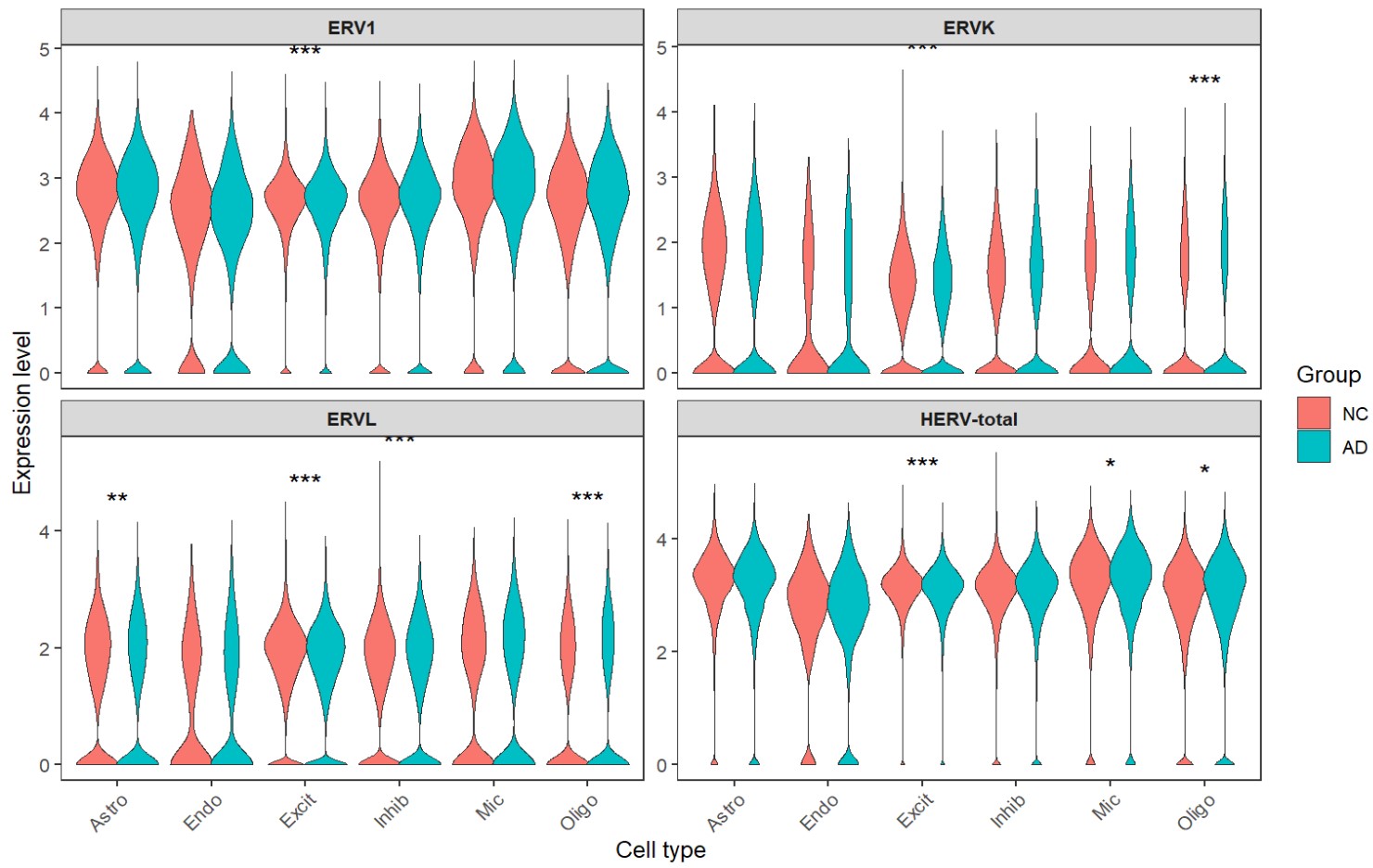
综上,hERV总体在全体细胞中并没有特别明显的重调(RVL/ERVK略低、ERV1略高,但效应量极小),如果按细胞类型拆开:
- Excit(兴奋性神经元):一致下调,且logFC相对较大
- Oligo(少突胶质细胞):一致上调
- Astro(星形胶质细胞):ERVL下调
- Mic(小胶质细胞):total轻微上升
- 其它细胞效应都较小,或由于细胞数过少而导致padj较大
step2:家族重调情况
DefaultAssay(seu) <- "HERV_family"
x <- GetAssayData(seu, assay = "HERV_family", slot = "data")
run_de_one_celltype <- function(seu, ct) {
if(ct!="All"){
obj <- subset(seu, subset = celltype == ct)
}else{
obj <- seu
}
DefaultAssay(obj) <- "HERV_family"
n_tab <- table(obj$group)
n_NC <- unname(n_tab["NC"])
n_AD <- unname(n_tab["AD"])
Idents(obj) <- factor(obj$group, levels = c("NC","AD"))
res <- FindMarkers(
object = obj,
ident.1 = "AD",
ident.2 = "NC",
test.use = "MAST",
latent.vars = c("orig.ident", "nCount_RNA", "percent_mito"),
min.pct = 0.05,
logfc.threshold = 0
)
res <- res %>%
tibble::rownames_to_column("subfamily") %>%
mutate(
celltype = ct,
n_NC = n_NC,
n_AD = n_AD
) %>%
relocate(celltype, n_NC, n_AD, subfamily)
res
}
celltypes_to_run <- c("Excit", "Oligo", "Astro", "Mic", "All")
res_list <- lapply(celltypes_to_run, function(ct) run_de_one_celltype(seu, ct))
res_list <- Filter(Negate(is.null), res_list)
res_subfamily <- bind_rows(res_list)
save(res_subfamily, file = "C:\\Users\\17185\\Desktop\\hERV_calc\\GSE157827\\my_data\\0303_1.RData")
write.csv(res_subfamily, file = "C:\\Users\\17185\\Desktop\\hERV_calc\\GSE157827\\my_res\\hERV_family.csv", row.names = F)
- Excit:广泛下调,包括很多HERVK家族
- Oligo:6个显著条目,HERVK下调,HERVH上调,部分与Excit方向相反
- Astro:只有2个显著,但HERVK(HML2)上调,与前两个不同
- Mic:只有1个显著且效应小
注意:有一些条目的logFC和pct相反,即平均表达量和表达比例高低不符,说明可能是在某个亚群中更特异表达
step3:细胞亚群中的家族重调情况
就从前面结果最显著的Excit细胞开始,先用常规方法把Excit细胞划分成更细的亚群
excit <- subset(seu, subset = celltype == "Excit")
DefaultAssay(excit) <- "RNA"
excit <- FindVariableFeatures(
excit,
selection.method = "vst",
nfeatures = 2000
)
excit <- ScaleData(
excit,
features = VariableFeatures(excit),
vars.to.regress = c("nFeature_RNA", "percent_mito")
)
excit <- RunPCA(
excit,
features = VariableFeatures(excit),
)
ElbowPlot(excit, ndims = 50)
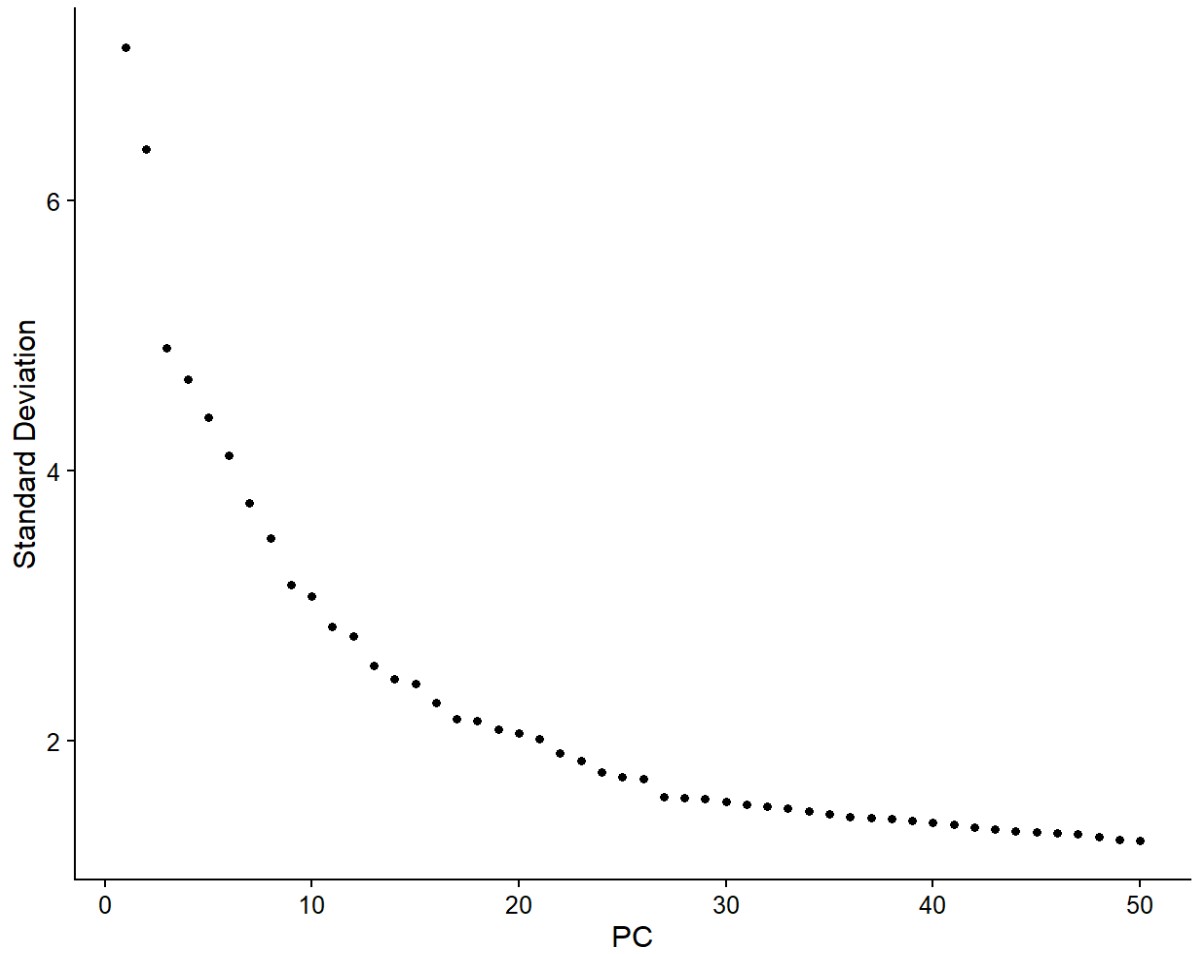
pc.num <- 1:30
excit <- RunUMAP(
excit,
dims = pc.num
)
DimPlot(excit, group.by = "group") | DimPlot(excit, split.by = "group")
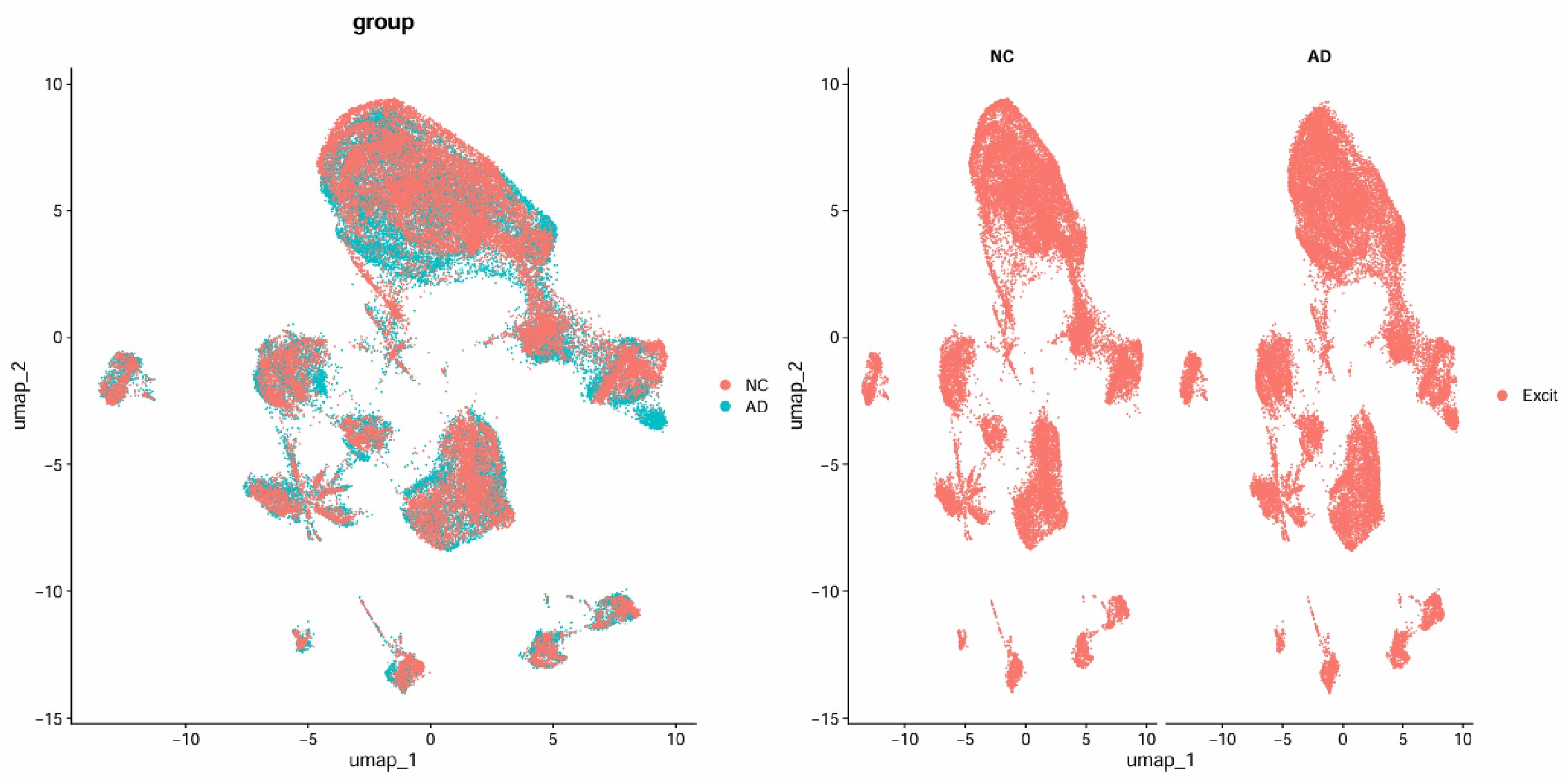
excit <- FindNeighbors(
excit,
dims = pc.num
)
excit <- FindClusters(
excit,
resolution = c(0.01,0.05,0.1,0.2,0.3,0.4,0.5)
)
clustree(excit@meta.data, prefix = "RNA_snn_res.")
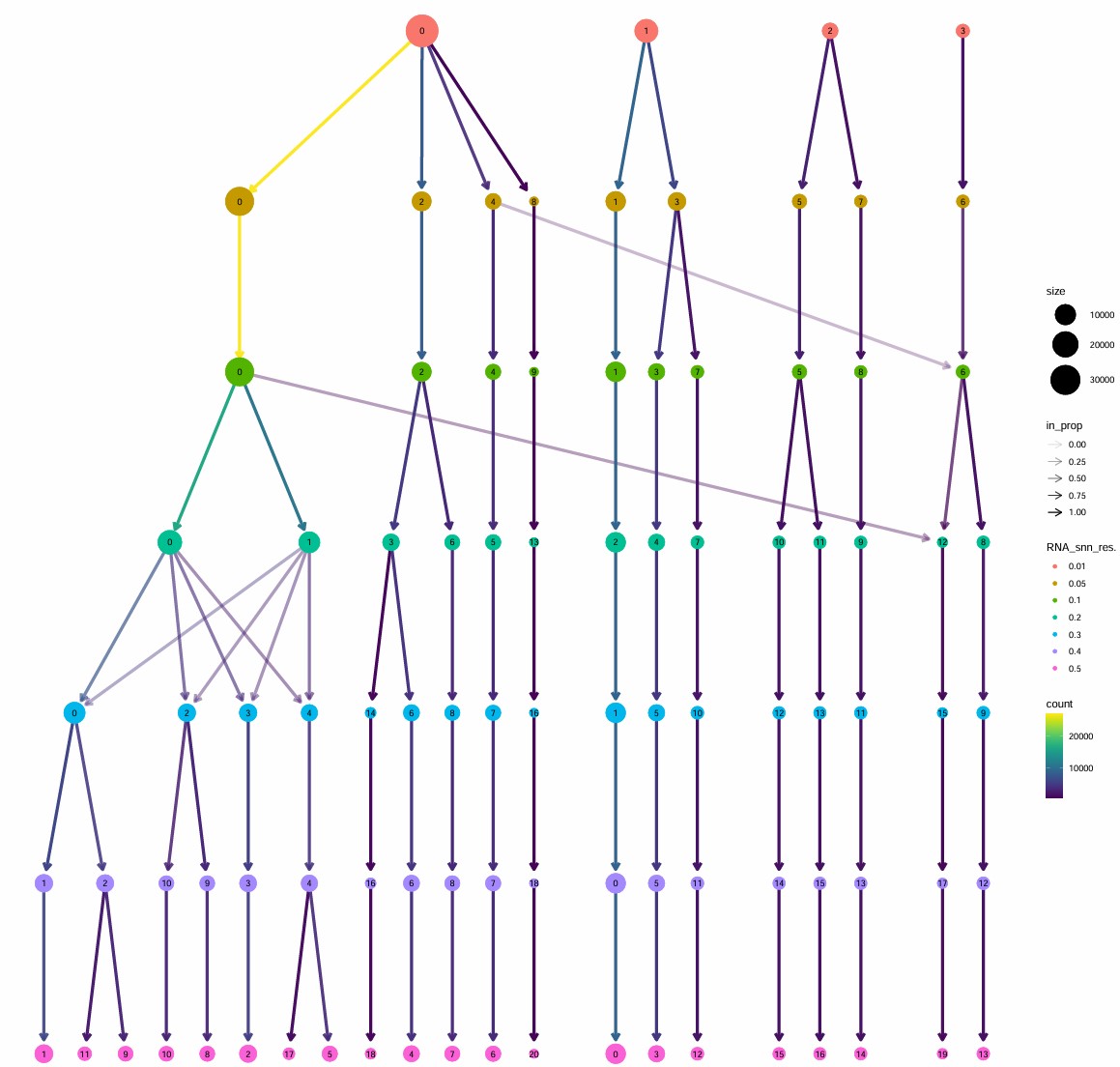
# 选res=0.3作为最终Excit子簇
excit$excit_subcluster <- excit$RNA_snn_res.0.3
Idents(excit) <- excit$excit_subcluster
DimPlot(excit, reduction = "umap", group.by = "excit_subcluster", label = TRUE)
# 保存数据
saveRDS(
excit,
file = "C:\\Users\\17185\\Desktop\\hERV_calc\\GSE157827\\my_data\\GSE157827_excit.rds",
compress = "xz"
)
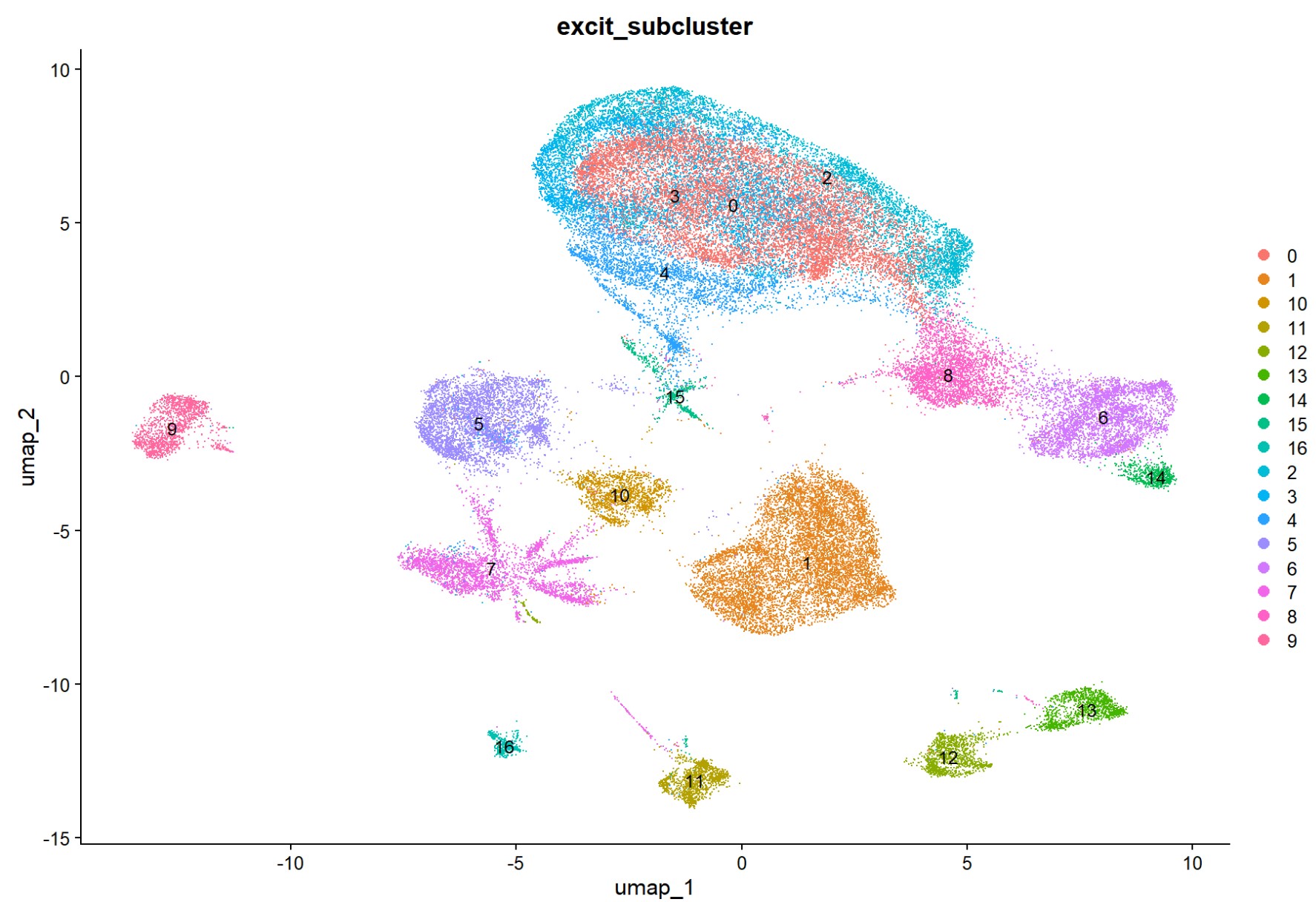
> table(excit$excit_subcluster)
0 1 10 11 12 13 14 15 16 2
11094 8831 1512 1290 1280 1264 648 545 418 5740
3 4 5 6 7 8 9
5663 4470 4359 4166 3320 3150 1515
> prop.table(table(excit$group, excit$excit_subcluster), margin=2)
0 1 10 11 12
NC 0.65494862 0.47333258 0.41600529 0.47751938 0.41718750
AD 0.34505138 0.52666742 0.58399471 0.52248062 0.58281250
13 14 15 16 2
NC 0.44462025 0.01080247 0.47522936 0.39712919 0.46498258
AD 0.55537975 0.98919753 0.52477064 0.60287081 0.53501742
3 4 5 6 7
NC 0.33586438 0.36510067 0.40467997 0.55592895 0.44548193
AD 0.66413562 0.63489933 0.59532003 0.44407105 0.55451807
8 9
NC 0.50507937 0.43696370
AD 0.49492063 0.56303630
总体来看其实每个亚群内AD和NC的比例差的不是很多,只有3/4/5/10/12/13这几个比较偏AD,0/6比较偏NC
- 14这个比较特殊,看了下样本分布,发现几乎全部来自AD5样本,感觉属于异常细胞
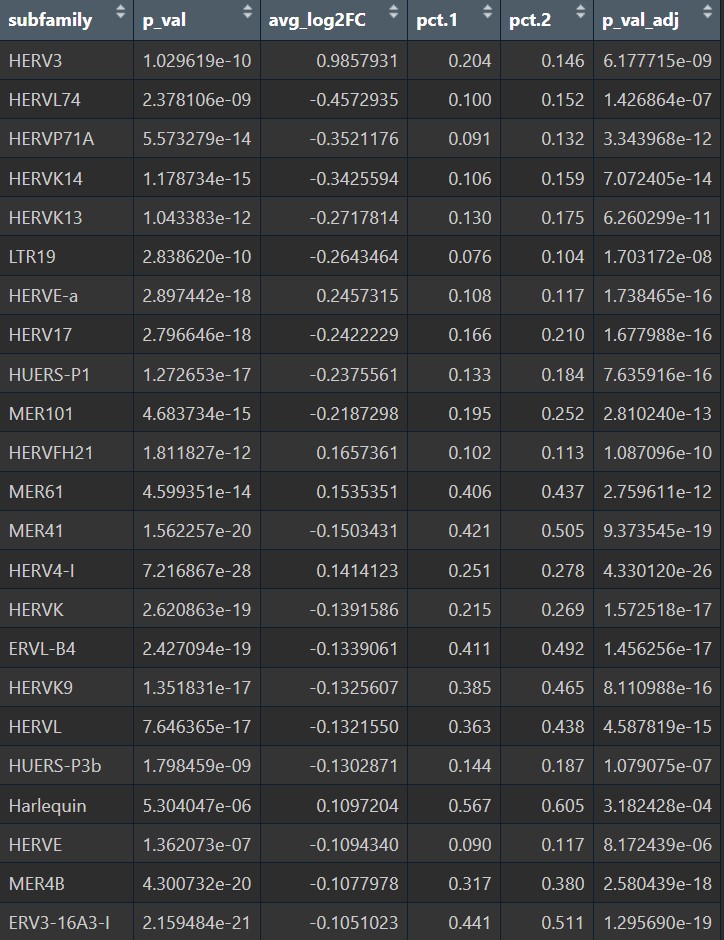
# 选出表达差异较显著的
res_subfamily <- read.csv("C:\\Users\\17185\\Desktop\\hERV_calc\\GSE157827\\my_res\\hERV_family.csv")
res_excit_focus <- res_subfamily %>%
filter(
celltype == "Excit",
!is.na(p_val_adj),
p_val_adj < 0.05,
abs(avg_log2FC) >= 0.1,
pmax(pct.1, pct.2) >= 0.10, # 至少一组有一定检出
pmin(pct.1, pct.2) >= 0.02 # 另一组不要过于稀疏
) %>%
arrange(desc(abs(avg_log2FC)), p_val_adj)
down_subfam <- res_excit_focus %>% # 下调
filter(avg_log2FC < 0) %>%
arrange(avg_log2FC) %>%
pull(subfamily)
up_subfam <- res_excit_focus %>% # 上调
filter(avg_log2FC > 0) %>%
arrange(desc(avg_log2FC)) %>%
pull(subfamily)
plot_subfam <- c(head(down_subfam, 6), head(up_subfam, 6))
Idents(excit) <- excit$excit_subcluster
DefaultAssay(excit) <- "HERV_family"
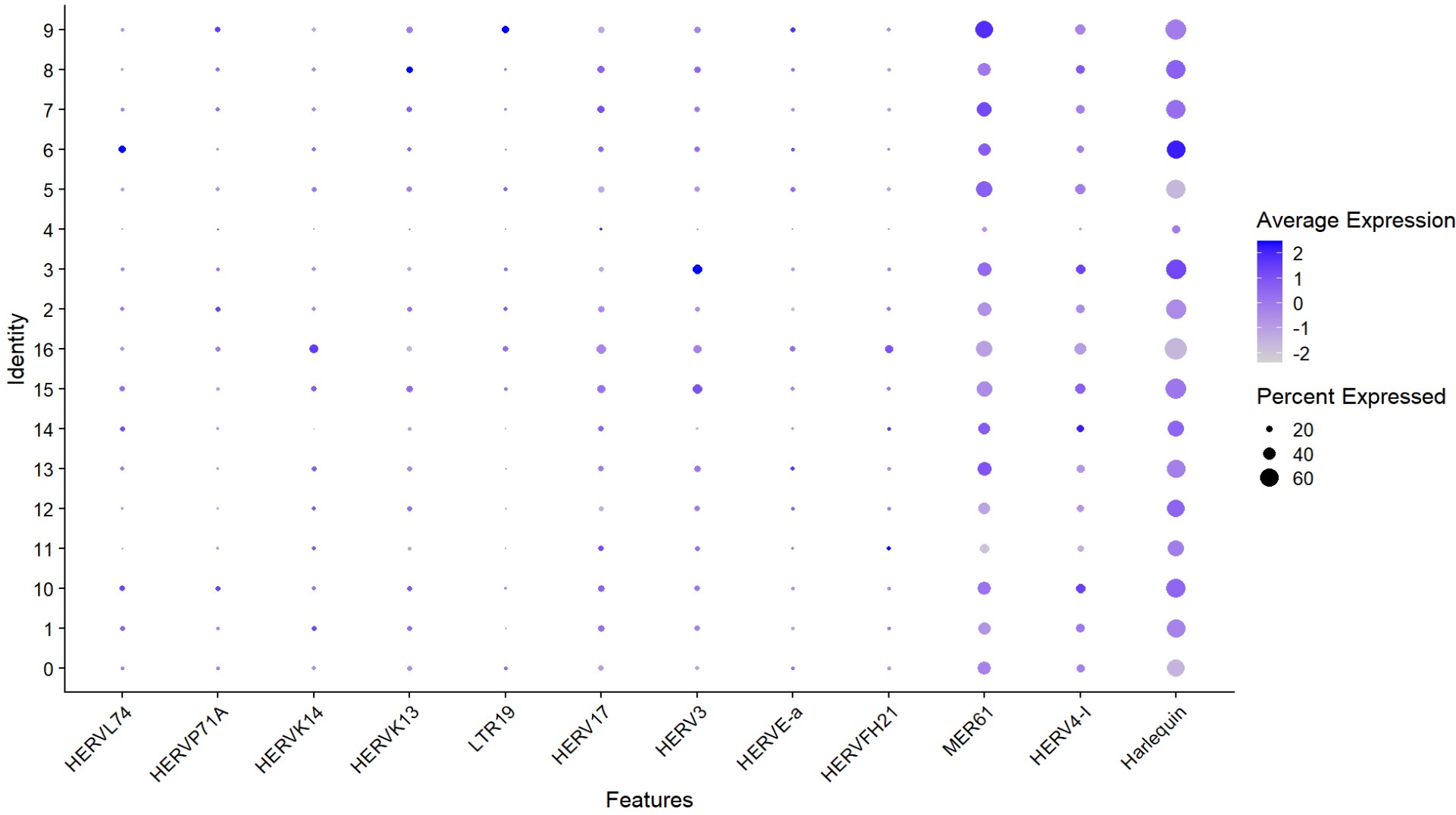
DotPlot(
excit,
features = plot_subfam,
group.by = "excit_subcluster",
assay = "HERV_family"
) + RotatedAxis()
plot_subfam <- c(down_subfam, up_subfam)
avg_mat <- AverageExpression(
excit,
assays = "HERV_family",
features = plot_subfam,
group.by = "excit_subcluster",
slot = "data"
)$HERV_family
avg_mat_z <- t(scale(t(avg_mat)))
avg_mat_z[is.na(avg_mat_z)] <- 0
pheatmap::pheatmap(
avg_mat_z,
cluster_rows = TRUE,
cluster_cols = FALSE,
fontsize_row = 10,
fontsize_col = 10
)
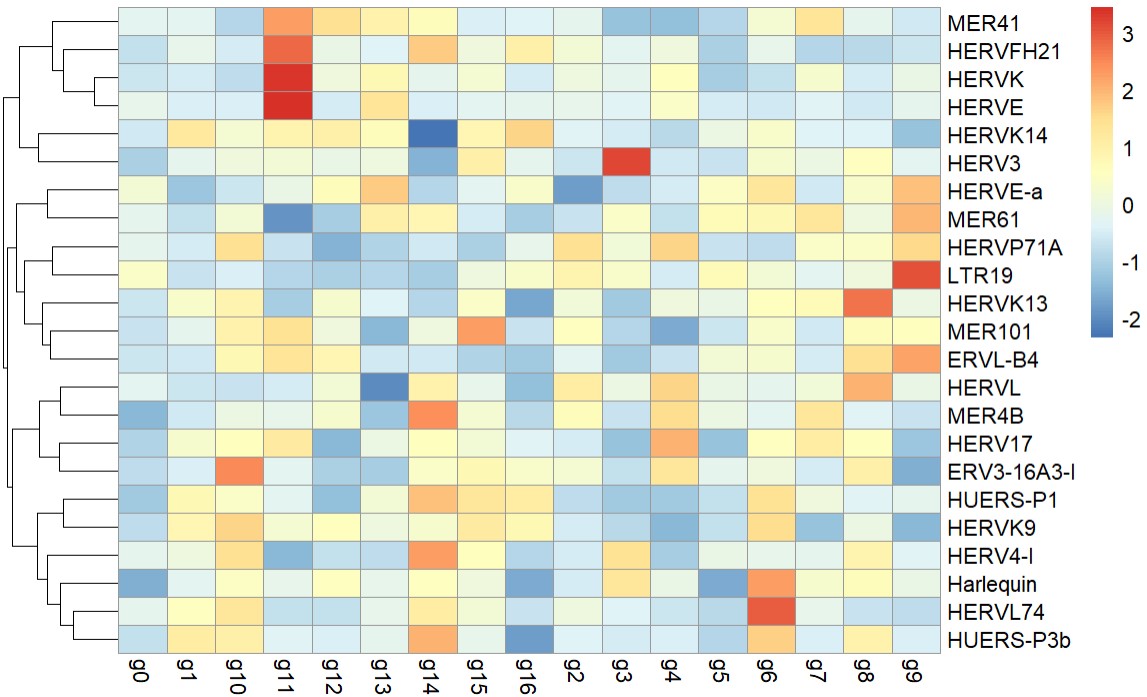
仍把上调和下调的hERV家族分别聚合,看聚合后结果在亚群中的分布?
herv_data <- GetAssayData(excit, assay = "HERV_family", slot = "data")
excit$herv_down_score <- Matrix::colMeans(herv_data[down_subfam, , drop = FALSE])
excit$herv_up_score <- Matrix::colMeans(herv_data[up_subfam, , drop = FALSE])
excit$herv_contrast_score <- excit$herv_up_score - excit$herv_down_score
FeaturePlot(
excit,
features = c("herv_down_score", "herv_up_score", "herv_contrast_score"),
reduction = "umap",
order = TRUE,
ncol = 3,
min.cutoff = "q05",
max.cutoff = "q95"
)
ad_global <- mean(excit$group == "AD")
cluster_summary <- excit@meta.data %>% # 统计各亚群的AD比例和hERV分数
group_by(excit_subcluster) %>%
summarise(
n = n(),
ad_prop = mean(group == "AD"),
ad_enrich = ad_prop - ad_global,
herv_down_mean = mean(herv_down_score, na.rm = TRUE),
herv_up_mean = mean(herv_up_score, na.rm = TRUE),
herv_contrast_mean = mean(herv_contrast_score, na.rm = TRUE),
.groups = "drop"
) %>%
arrange(desc(ad_prop))
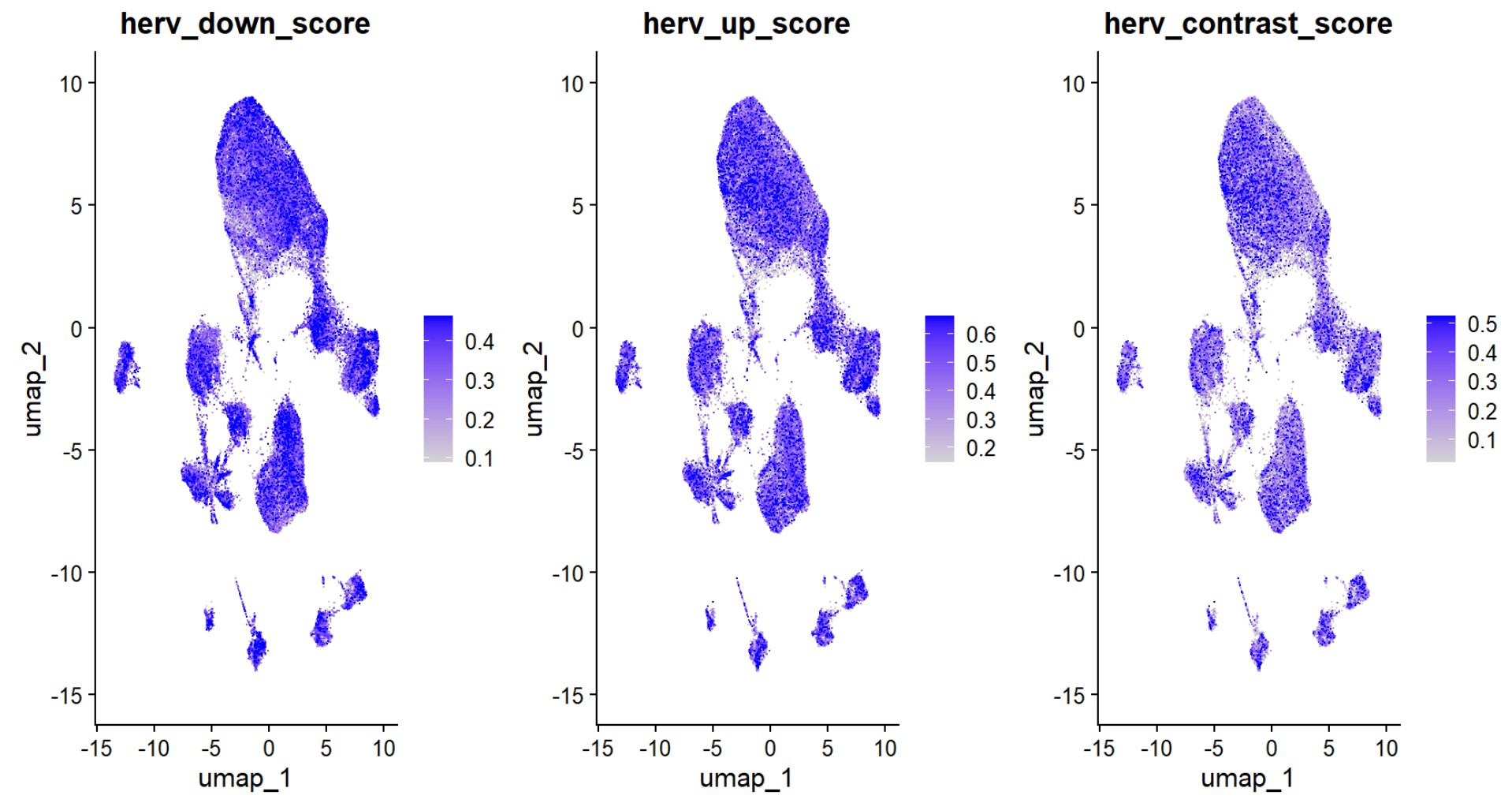
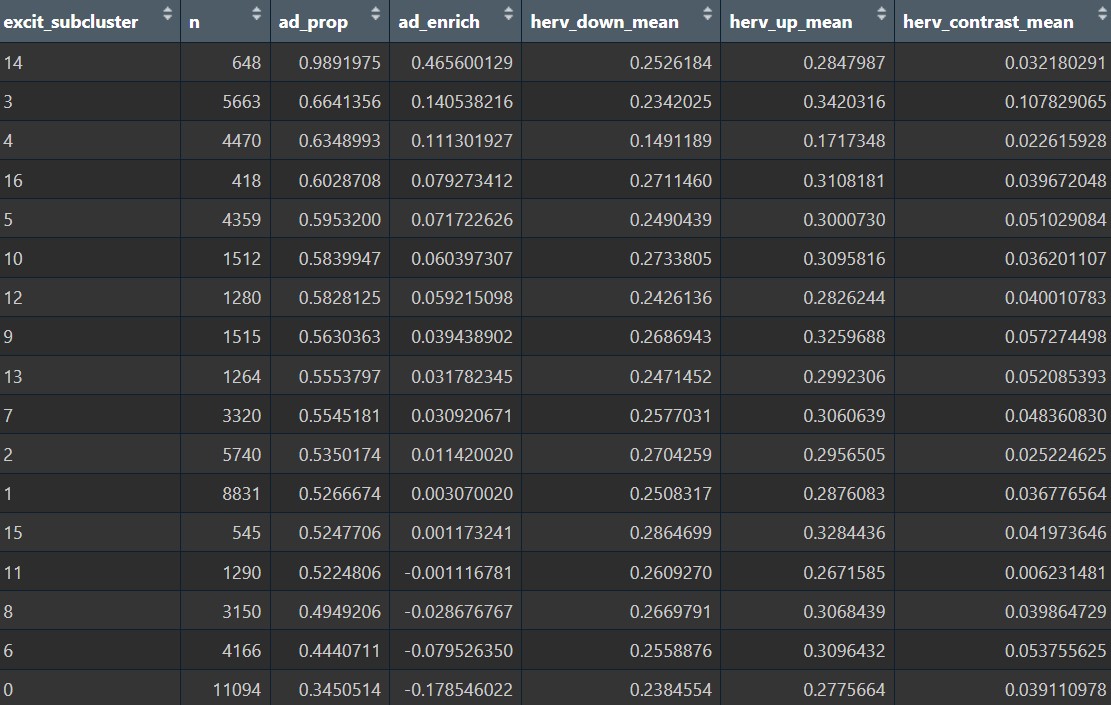
可以看到,hERV分数在整个Excit上分布比较广、比较连续,没有哪几个亚群突出表达hERV,且AD状态与hERV分数相关性较弱。总的来说,把上调/下调的HERV家族简单平均成一个总分,并不能很好地区分Excit的AD-rich/NC-rich亚群
- 不同hERV家族的空间分布和与AD的关系并不一致
- 是否有某些具体hERV家族对应某些具体亚群?(看单个或少数几个hERV家族的局部模式,而不是总分)
- HERV3在cluster 3(AD-rich)很突出
- hERVK/E在cluster 11很突出,但cluster 11的AD比例接近Excit平均水平,没有显著偏离
- HERVL74在cluster 6(NC-rich)较突出
- MER41/HERVFH21/HERVK/HERVE:在某些亚群里更高,但不一定对应最AD-rich的群
哪些亚群更适合作为研究对象:
- 从
偏AD/NC的角度:- cluster 5:AD-rich、4k多个细胞、样本分布相对最均匀
- cluster 3/4:AD-rich、细胞数够但样本有一定偏倚
- cluster 0/6:NC-rich
- 从
hERV重调的角度:- cluster 3:HERV3
- cluster 6:HERVL74
- cluster 11:hERVK/E
这之后又是之前的gene~hERV表达做法了?只是只看某个hERV家族
文献阅读
一些与”HERV/AD”相关的文献
都是RNA-seq的
- Locus specific endogenous retroviral expression associated with Alzheimer’s disease
- 组织:Parietal cortex(顶叶皮层)/Anterior temporal lobe(前颞叶)/Lateral temporal lobe(侧颞叶)
- 方法:bulk RNA-seq+Telescope位点级分析
- 样本数:103AD+45NC
- Human endogenous retrovirus HERV-K(HML-2) RNA causes neurodegeneration through Toll-like receptors
- 组织:temporal cortex(颞叶皮层, TCX)+cerebellum(小脑),其中只有TCX中HERV-K有差异
- 方法:bulk RNA-seq+featureCounts家族级分析
- 样本数:82AD+94NC
- 两篇都主要说了HERVK与TLR8上调/相关,特别是HML2
- 位点级分析:698个hERV位点中,187个下调,511个上调;HERVK家族层面相关性不显著,部分位点和subfamily层面显著(HML1/2/3/4/5/6、HERVK11、HERVK11D和HERVKC4)
我使用的数据集:
- 组织:prefrontal cortex(前额皮层)
- 方法:snRNA-bulk+Stellarscope位点级分析
- 样本数:12AD+9NC
可以看看hERVK位点与免疫基因(TLR8、TLR2)的共表达情况?聚合到样本层面再计算相关性?TLR8相关的hERVK位点是不是主要出现在Mic/Endo或某些反应性亚群里?
Retrotransposons unplugged
Retrotransposons unplugged: Rewiring the nervous system and wreaking havoc
LTR型逆转座元件和ERV相关序列(retroTEs):占人类基因组的8%-10%,其中很多位点虽然已经退化,通常不再活跃转座,但仍然能够被转录,甚至产生蛋白,从而影响细胞功能
- 在神经系统发育、髓鞘形成、突触可塑性和记忆等正常生理中发挥作用
- 在老化和神经退行性疾病中又可能通过炎症、蛋白毒性和细胞间传播等机制造成损伤
domesticated retroTE genes:一部分retroTE在进化中被宿主驯化成了新基因,这些基因虽然已经不再是完整病毒,但保留了某些关键结构域,比如Gag-like capsid domain、RNA结合结构域、蛋白酶结构域,因此可能仍具有“病毒样”的生物学能力。它们可能参与了神经系统中的一些复杂过程,比如细胞间通讯、记忆形成、免疫调节,当表达失衡时,这些机制就会变成病理来源
- Arc:经典的神经元立早基因/即刻早期基因(immediate early genes)——在受到外界刺激后迅速并且短暂激活的基因,不需要合成新蛋白。在序列和结构上都类似逆转录病毒的Gag基因,能够自发形成类似病毒衣壳的结构,并把Arc mRNA封装进细胞外囊泡,在细胞之间转运,从而介导一种新型的细胞间突触可塑性。宿主其实已经“借用”了逆转座元件的病毒样能力来完成高级神经功能
- PNMA2:在神经元中表达,可以自组装形成无包膜衣壳,并且高度免疫原性,可能在大脑中参与关键功能。小鼠被注射PNMA2衣壳后会产生强烈自身抗体反应,并出现学习记忆缺陷。retroTE蛋白可能以病毒样颗粒的形式参与神经系统生物学和免疫相关病理
- PEG10/RTL(Mart家族):这类gag-like基因与行为、神经功能、微胶质相关反应以及神经发育异常有关
- 缺乏PEG11/RTL1的小鼠会出现焦虑行为增强以及学习和记忆障碍
- RTL5和RTL6在小胶质细胞中表达,可能调控大脑中的先天免疫反应
- RTL4是小胶质细胞分泌的因子,与自闭症有关
- RTL8家族与行为调控(社交响应)有关
- PEG10的表达对胎盘发育至关重要,其水平升高可能与癌症发生有关,其异常积累可能直接导致神经疾病,还能形成并释放病毒样颗粒,并可能介导核酸的细胞间转运
病毒样颗粒(virus-like particles, VLPs):由病毒单一或多个结构蛋白自行组装形成的空心蛋白颗粒,其保留了病毒抗原的天然构象但不含遗传物质,不具备传染性与致病性
retroTE不仅和神经元有关,也与发育相关
- 源自retroTE的RNLTR12-int(RetroMyelin)对髓鞘形成至关重要,可调节髓鞘碱性蛋白的表达
-
HERV-K相关蛋白在胚胎和干细胞中也被观察到,与先天免疫调控因子的上调有关
囊胚形成过程中观察到VLP,但目前还不清楚这些颗粒具体如何介导HERV-K相关的基因表达变化
retroTE产生的衣壳蛋白在结构上和病毒衣壳相似,它们也许同样可以承担稳定包裹并远距离转运RNA/蛋白、提高信号传递效率的功能。总的来说,retroTE为动物进化提供了一套现成的分子运输与通信模块,而神经系统和发育过程可能反复利用了这套模块
- 完整env基因的表达可能促进VLP被远端细胞摄取
- 线虫中的Ty3retroTEcer1,可能通过VLP参与RNA介导的跨代学习记忆
- 斑马鱼中的Bik-1/Bik-2,这两个可能编码形成capsid的Gag蛋白,参与中胚层和神经嵴发育
retroTE与神经退行性疾病:
- 逆转录病毒的急性感染可通过炎症信号的启动、诱导蛋白质毒性和DNA损伤,引发神经退行性疾病并加重神经系统疾病
- HIV感染可能导致HIV相关神经认知障碍,其特征是认知障碍和脑部灰质丧失,类似于AD;也可能引起一种类似ALS的病症,表现为运动神经元退化,与ALS不同的是,该病对抗逆转录病毒治疗有反应
- 疱疹病毒与AD有关,接种带状疱疹疫苗的人,换AD的可能性较低
- 外源性病毒感染可能干扰由驯化或内源性retroTE介导的正常功能:Arc与疱疹病毒(HSV-1)的表达和成熟有关,HSV-1的衣壳蛋白似乎对病毒在细胞内摄取后增殖至关重要
- 随年龄增长,retroTE会去抑制而表达升高,这与ALS、FTD、AD等疾病相关
- 疱疹病毒阳性组织的AD脑组织中LINE表达更高,而且体外可被抗病毒处理部分逆转
- 在部分ALS患者组织中反复观察到HERV-K表达升高,HERV-K env的过度表达直接导致小鼠运动和行为功能的神经退行性下降,使用RT inhibitor后AD症状有一定减缓
- EBV病毒会使多发性硬化症(MS)风险增加30多倍,HERV-W表达的激活可能是疾病的诱因
retroTE影响神经系统的机制:
- 神经炎症:retroTE表达后会产生RNA、DNA和蛋白产物,这些分子可被MDA5、RIG-I、AIM2、cGAS/STING、TLR7/8、TLR4等免疫感受器识别,诱导先天免疫和后续炎症反应
- 表达HERV-K的细胞死亡会释放RNA进入细胞外空间,从而激活TLR7/8
- tau诱导的retroTE去抑制可产生dsRNA,并激活MDA5通路
- 逆转录酶抑制剂可缓解AD症状,但不清楚是直接调控逆转录元件,还是通过靶向外效应,抑制潜伏的逆转录病毒感染
- 细胞内毒性和信号扰动:某些retroTE的gag或env蛋白本身就存在细胞毒性,或者改变神经元基因表达和蛋白稳态
- PEG10在Angelmansyndrome和ALS中升高,影响树突/轴突重塑相关基因表达
- PEG10和RTL8改变应激颗粒动态,导致蛋白质稳态变化,引发神经功能障碍
- 发育中小鼠大脑中PEG10的过度表达会改变神经元迁移
- PEG10改变基因表达的分子机制和引发神经退行的机制尚不明确
- VLP介导的细胞间传播:retroTE或其驯化基因形成的VLP可能像“运输工具”一样,在细胞之间传播RNA或错误折叠蛋白,例如蛋白质聚集体(protein aggregates),从而促进病理扩散
- VLPs可能促进蛋白质聚集体的形成和传播:小鼠retroTE IAP(intracisternal A-particles) Gag编码的衣壳蛋白在tau转基因小鼠的大脑中也增多,并与记忆障碍相关
- Arc与AD病理和突触功能障碍有关,可能促进tau被装入EV/VLP并在细胞间转运,Arc缺失会减少tau的传播
- HERV-K和果蝇MDG-ERV既能触发TDP-43聚集,也促进TDP-43病理的细胞间传播
- Arc和PEG10衍生的VLP含有mRNA,可将特定遗传物质从一个细胞转移到另一个细胞,可能通过引入破坏细胞身份或改变细胞行为的RNA,改变神经元功能,但具体包裹的mRNA尚不明确
未来展望:
- 目前retroTE研究最大的瓶颈还是技术层面,因为这类序列高度重复、不同位点之间很相似,传统短读长RNA-seq很难准确比对,所以过去很多常规分析流程其实会直接把retroTE信号排除掉。现在虽然已经有一些新的分析工具,但不同流程之间结果仍不完全一致,还缺少一个统一、可靠的retroTE图谱
- 长读长测序有希望更好地区分这些相似序列,不过目前成本和测序深度仍是限制
- retroTE蛋白产物的鉴定也很困难,因为标准蛋白组数据库往往不收录这些蛋白,需要更好的测序、比对、注释和蛋白鉴定工具
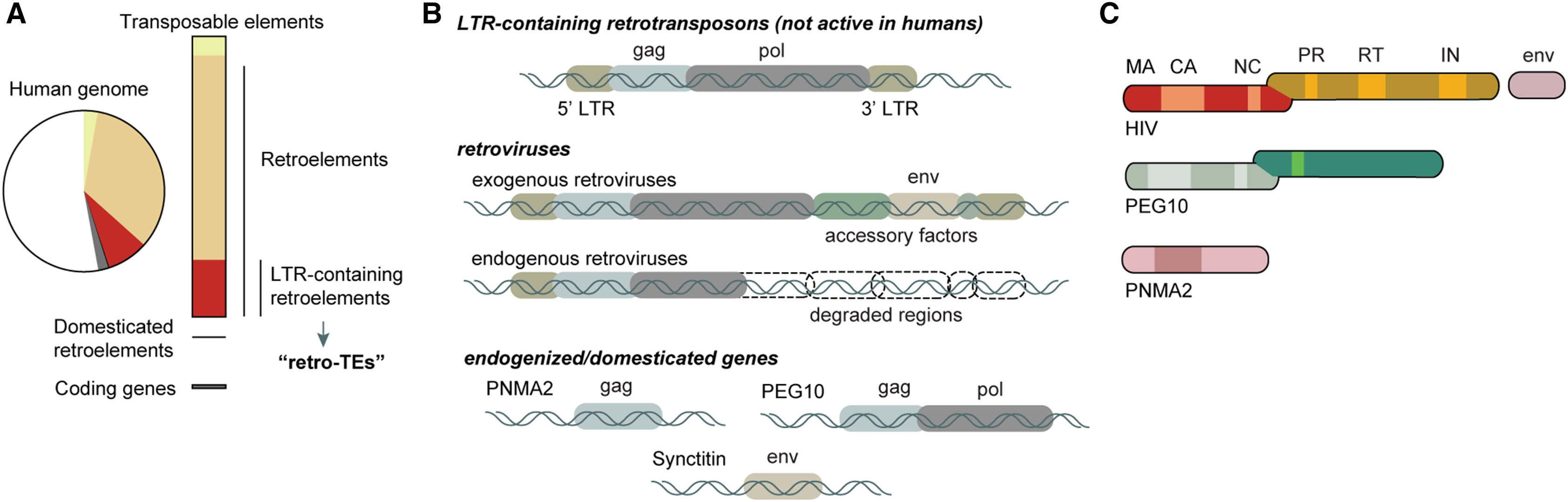
- 逆转座子和ERV保留更多病毒样结构,但但人类基因组中的多数ERV位点已经退化(被宿主“驯化”后的ERV衍生基因),缺少完整复制能力和LTR调控,只保留部分开放阅读框,并改用宿主自身的启动子和增强子调控
- 这些驯化基因的蛋白仍可保留类似gag、gag-pol或env的结构特征
- gag:编码病毒的核心蛋白(核衣壳蛋白、内膜蛋白、衣壳蛋白),Arc、PNMA2、PEG10等之所以常被说成有“病毒样”特征,核心就是它们保留了类似gag的成壳能力
- MA(matrix):基质层,靠近包膜内侧,主要帮助颗粒装配和定位
- CA(capsid):衣壳,病毒样颗粒最典型的外壳结构
- NC(nucleocapsid):核衣壳,主要负责结合RNA,也就是更贴近遗传物质的那一层
- pol:编码病毒复制相关的酶
- PR(protease):蛋白酶,负责把长的前体蛋白切开,变成成熟的功能蛋白
- RT(reverse transcriptase):逆转录酶,负责把RNA反转录成DNA
- IN(integrase):整合酶,负责把新生成的DNA整合到宿主基因组里
- gag-pol:一个蛋白如果像gag-pol,往往表示它既带一些结构特征,又带一些酶相关特征
- env:编码病毒包膜糖蛋白,ERV和一般LTR逆转座子的一个重要区别,就是ERV还有env基因,这有助于病毒样颗粒在细胞之间传播
- gag:编码病毒的核心蛋白(核衣壳蛋白、内膜蛋白、衣壳蛋白),Arc、PNMA2、PEG10等之所以常被说成有“病毒样”特征,核心就是它们保留了类似gag的成壳能力
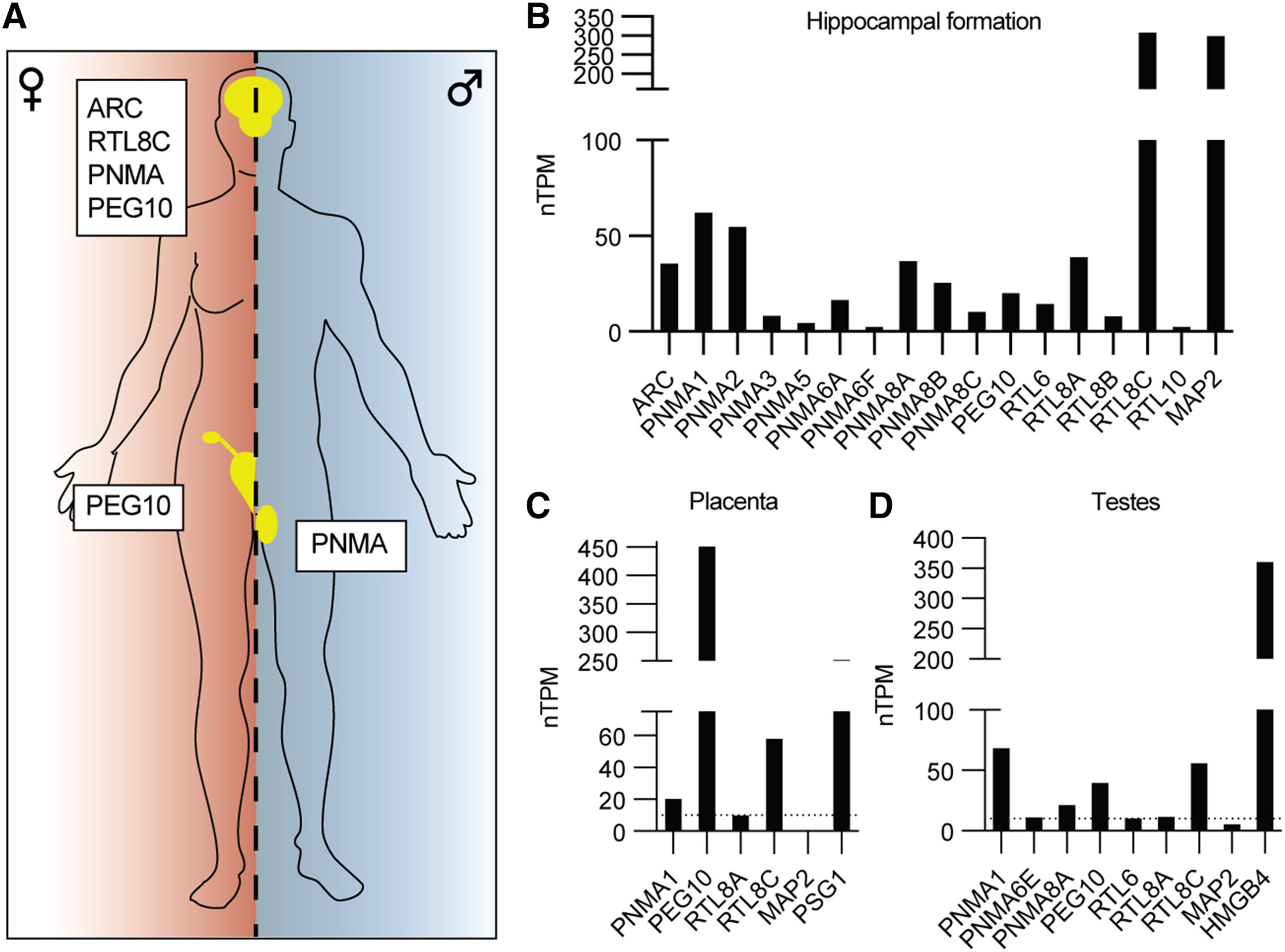
- 海马体/胎盘/睾丸中ERV相关基因的mRNA表达水平
- 参考基因:MAP2-神经组织,PSG1/HMGB4-胎盘/睾丸
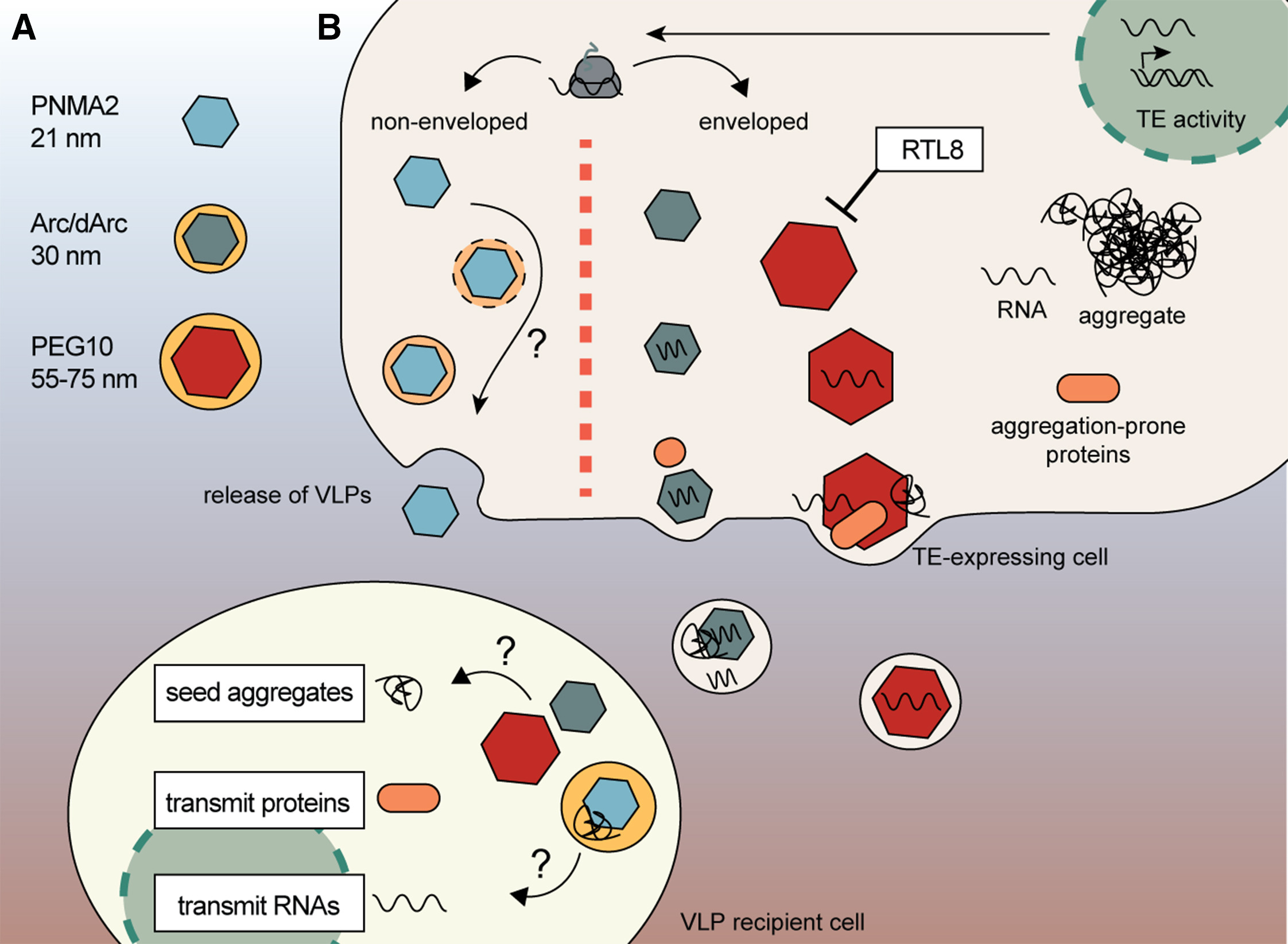
- VLP产物:PNMA2是无包膜的,而Arc和PEG10外面还有一层类似细胞外囊泡的脂质膜
- VLP在细胞中形成并释放,里面有时装的是RNA,有时也会同时装蛋白和RNA;进入受体细胞后,可以递送内容物,从而改变受体细胞行为
- 像Arc这样的VLP甚至可能装载错误折叠蛋白,例如tau,并在细胞间传播,推动病理缓慢扩散
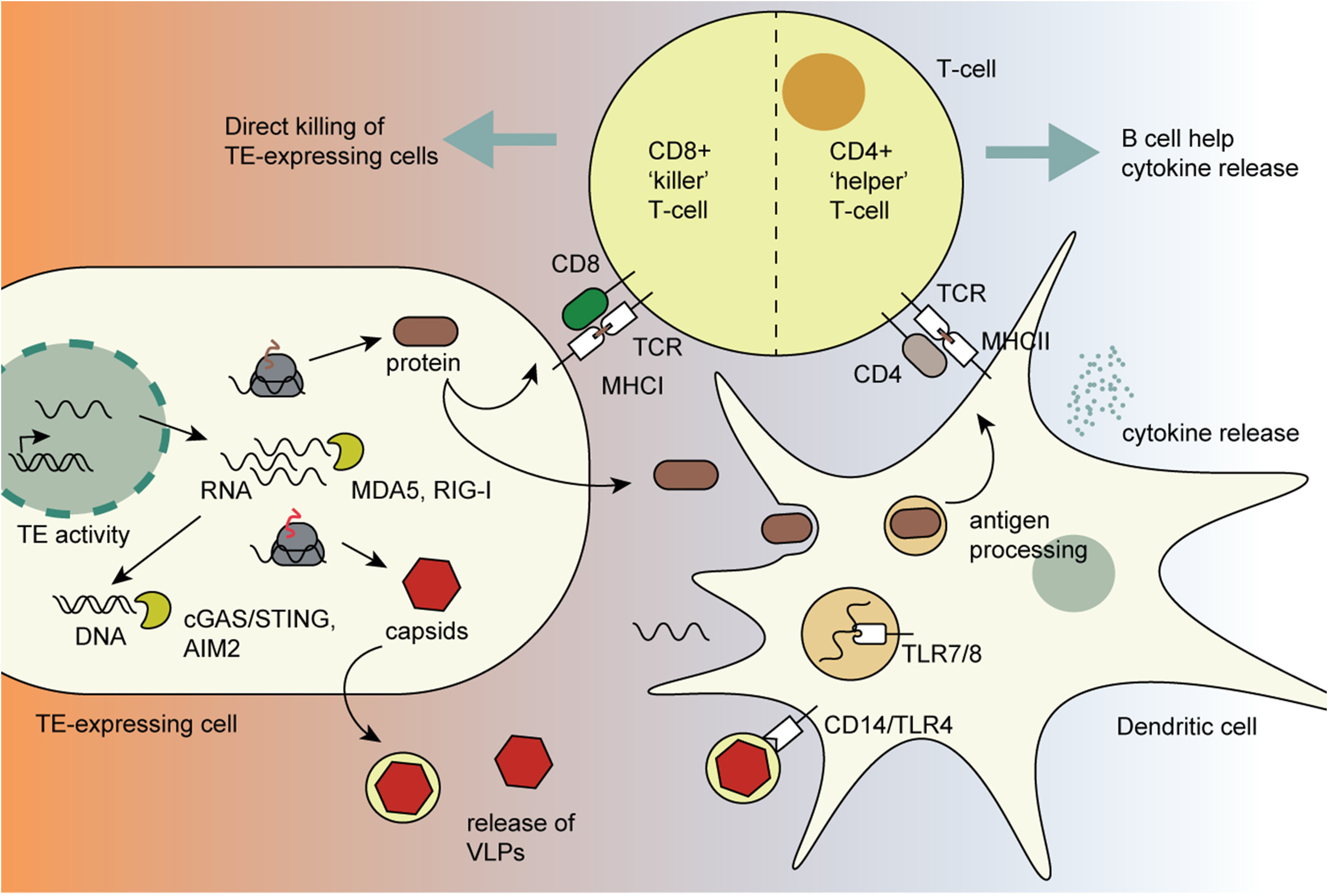
- retroTE转录出的RNA、逆转录产生的DNA会被先天免疫感受器识别,从而启动先天免疫
- 其蛋白产物一部分可直接经MHCI呈递,另一部分可被抗原呈递细胞摄取后经MHCII呈递给T细胞
- capsid蛋白还能组装成VLP,被CD14、TLR4等途径感知,诱导抗原呈递细胞释放炎症性细胞因子
- 最终,CD4和CD8T细胞都可能被激活,形成更强的适应性免疫反应
图1能不能做类似的
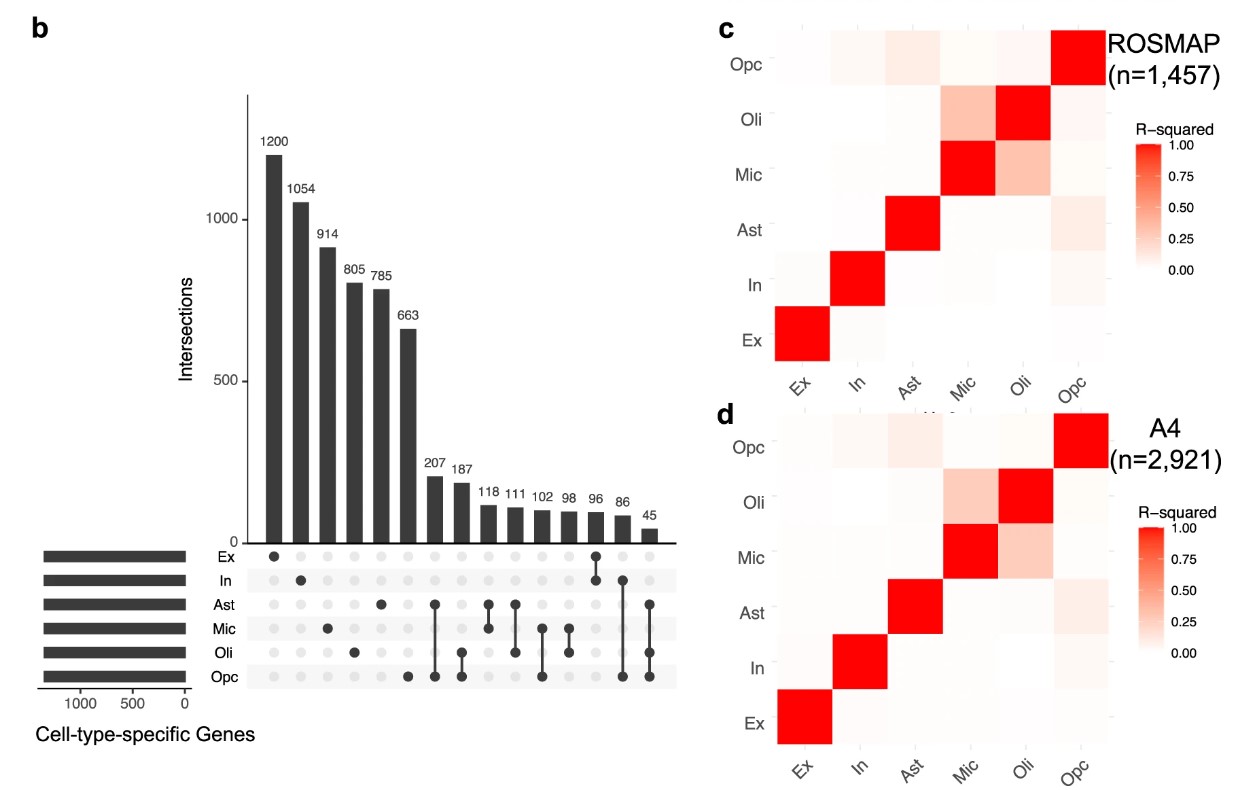
取六大细胞类型各自特异表达最高的前10%基因作为细胞特异性表达基因集
- 不同细胞类型特异基因集之间重叠情况的UpSetR图
- 不同细胞类型ADPRS之间相关性的相关矩阵,看各细胞类型的重调模式彼此像不像
ADPRS:AD polygenic risk scores,多基因AD风险评分,指基因i在细胞类型j的平均表达,占它在所有6类细胞平均表达总和的比例(相当于排除“普遍高表达”的影响)。然后把每个细胞类型里特异性最高的前10%基因定义为该细胞类型的基因集,结合GWAS,将每个样本的变异映射到这些基因集附近,最后得到每个样本的6个细胞类型特异性ADPRS(根据PRS/GWAS综合得到),图1c/d的相关矩阵是把这些分数两两做相关,看“某个体在Mic上AD风险高的时候,是否在Oli上也高”
自己画的话,要明确画的是“细胞类型特异hERV”还是“AD相关hERV”,作者的就是细胞类型特异,是比较每个细胞类型中基因的表达差异,这个思路可以直接借鉴到hERV里面,只是我们的hERV没有GWAS,只能用基因表达量;如果想探究每个细胞类型中AD导致的hERV重调模式是否相似,就用前面差异表达分析得到的每个细胞类型的显著hERV家族,画重叠关系,然后热图就用每个细胞类型每个hERV家族的logFC代替
- “细胞类型特异hERV”不反映AD带来的影响,原文似乎是只在AD样本中进行的(至少不是AD-NC对比进行的)
assay_use <- "HERV_family"
slot_use <- "data" # 使用标准化后的data
group_col <- "celltype"
# reference_group <- "NC" # 用哪种细胞画
reference_group <- NULL
min_pct_cells_express <- 0.01 # 至少在全部参考细胞中1%细胞表达
top_n <- 10 # 细胞类型特异性最显著的前10个基因(论文中为前10%)
if (!is.null(reference_group)) {
cells_use <- rownames(seu@meta.data)[seu@meta.data$group == reference_group]
} else {
cells_use <- colnames(seu)
}
meta_use <- seu@meta.data[cells_use, , drop = FALSE]
DefaultAssay(seu) <- assay_use
expr_mat <- GetAssayData(seu, assay = assay_use, slot = slot_use)[, cells_use, drop = FALSE]
# 过滤稀疏feature
detect_frac <- Matrix::rowMeans(expr_mat > 0)
expr_mat <- expr_mat[detect_frac >= min_pct_cells_express, , drop = FALSE]
# 每个细胞类型的平均表达,行名是
celltype_vec <- meta_use[[group_col]]
names(celltype_vec) <- rownames(meta_use)
celltypes <- sort(unique(as.character(celltype_vec)))
avg_expr_list <- lapply(celltypes, function(ct) {
ct_cells <- names(celltype_vec)[celltype_vec == ct]
if (length(ct_cells) == 1) {
as.numeric(expr_mat[, ct_cells, drop = FALSE])
} else {
Matrix::rowMeans(expr_mat[, ct_cells, drop = FALSE])
}
})
avg_expr_mat <- do.call(cbind, avg_expr_list)
rownames(avg_expr_mat) <- rownames(expr_mat)
colnames(avg_expr_mat) <- celltypes
# cell-type specificity score
row_sums <- rowSums(avg_expr_mat)
specificity_mat <- avg_expr_mat / ifelse(row_sums == 0, NA, row_sums)
specificity_mat[is.na(specificity_mat)] <- 0
mean_expr_overall <- rowMeans(avg_expr_mat)
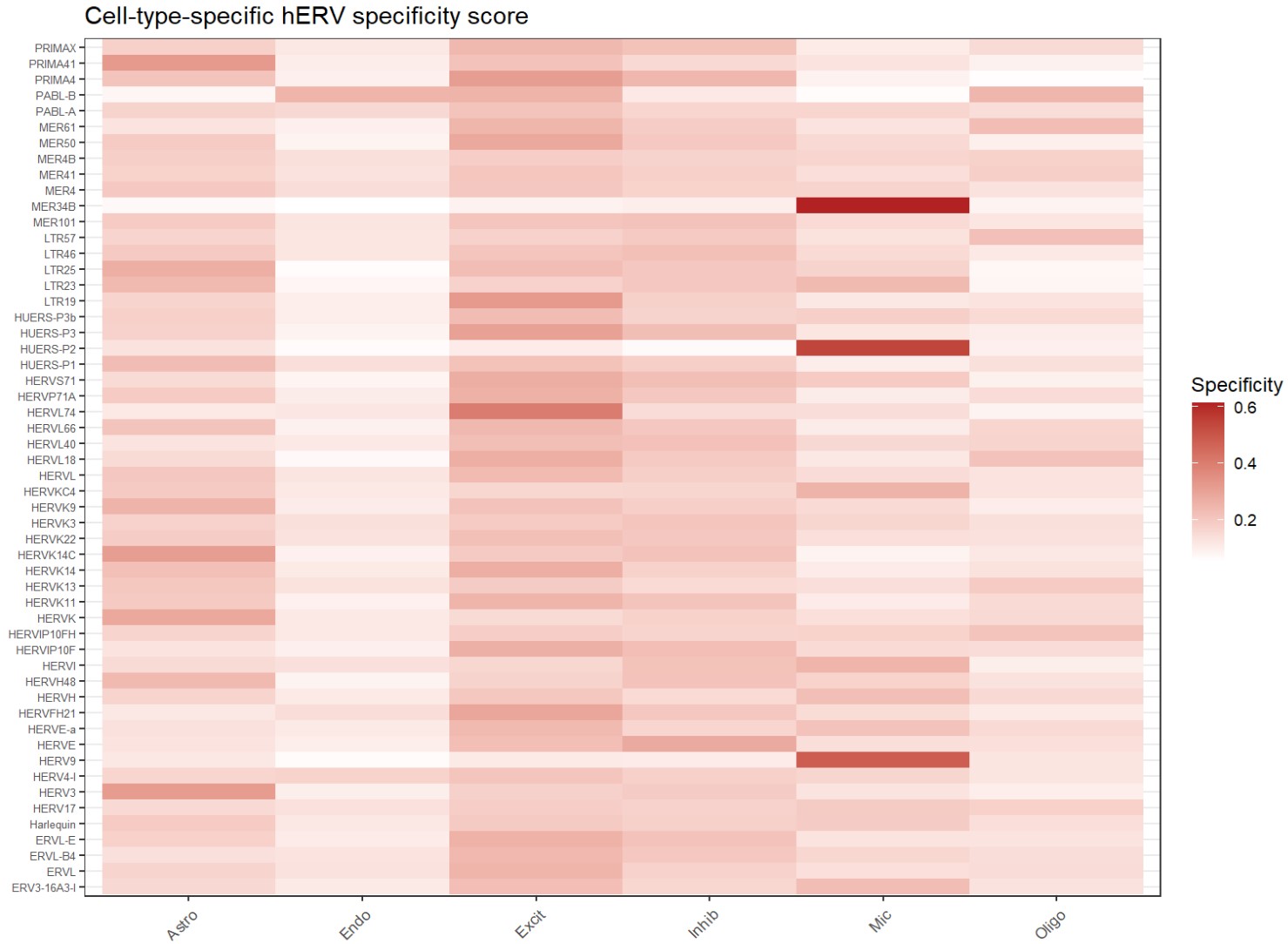
# specificity score 热图
df <- as.data.frame(specificity_mat) %>%
tibble::rownames_to_column("feature") %>%
tidyr::pivot_longer(-feature, names_to = "celltype", values_to = "score")
ggplot(df, aes(x = celltype, y = feature, fill = score)) +
geom_tile() +
scale_fill_gradient(low = "white", high = "firebrick") +
theme_bw() +
labs(title = "Cell-type-specific hERV specificity score", x = NULL, y = NULL, fill = "Specificity") +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
axis.text.y = element_text(size = 6)
)
# 为UpSet取每个细胞类型最特异的top families
top_features_by_ct <- lapply(colnames(specificity_mat), function(ct) {
score_vec <- specificity_mat[, ct]
names(sort(score_vec, decreasing = TRUE))[seq_len(min(top_n, length(score_vec)))]
})
names(top_features_by_ct) <- colnames(specificity_mat)
# 构建feature x celltype的membership矩阵
all_top_features <- sort(unique(unlist(top_features_by_ct)))
membership_df <- data.frame(feature = all_top_features, stringsAsFactors = FALSE)
for (ct in names(top_features_by_ct)) {
membership_df[[ct]] <- membership_df$feature %in% top_features_by_ct[[ct]]
}
# UpSet图
library(UpSetR)
ct_cols <- setdiff(colnames(membership_df), "feature")
upset_input <- membership_df[, ct_cols, drop = FALSE]
upset_input[] <- lapply(upset_input, as.integer)
UpSetR::upset(
upset_input,
sets = ct_cols,
keep.order = TRUE,
order.by = "freq",
mb.ratio = c(0.6, 0.4),
mainbar.y.label = "Intersection size",
sets.x.label = "Set size",
text.scale = 1.1
)
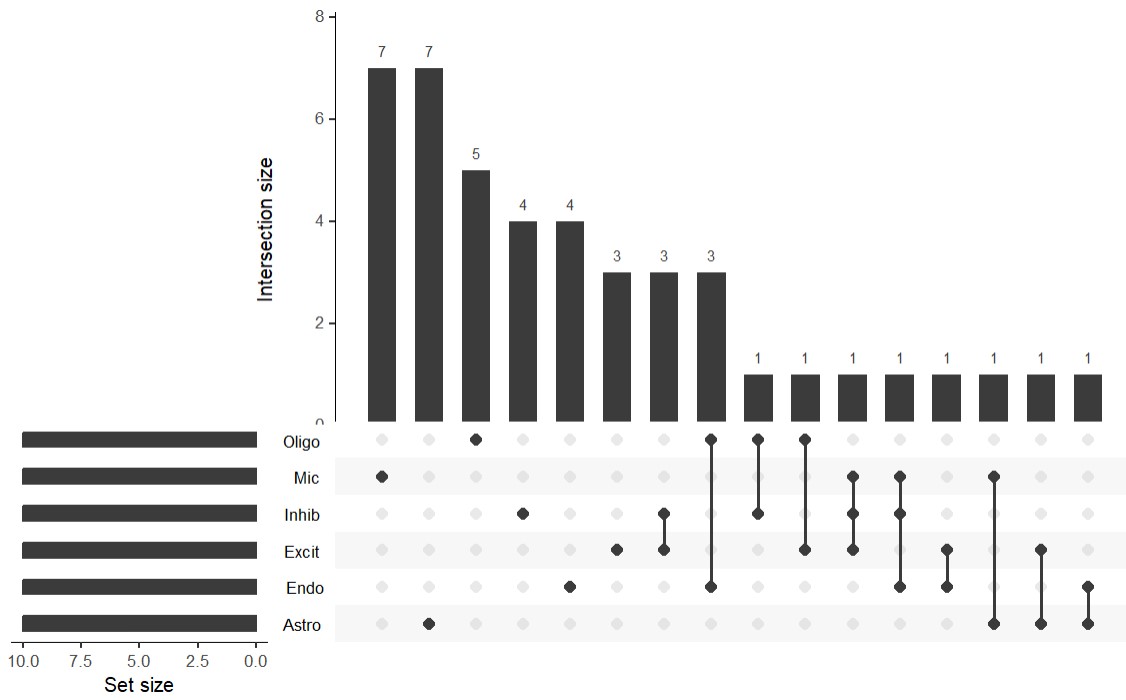
各细胞类型的tophERV家族既有明显的特异性,也有部分重叠,但独有成分占比较大
- 左下角Set size:每个细胞类型自己的集合大小,因为我在每个细胞类型中都取了前10各最特异的hERV家族,所以长度都是10
- 中下的点阵:当前这根柱子对应的是哪些细胞类型的交集
- 如果某一列只有Mic那一行是黑点,别的都是灰点:这一列代表的是只出现在Mic的hERV家族
- 如果某一列Excit和Inhib两行是黑点,并且中间有线连起来:这一列代表的是同时出现在Excit和Inhib的hERV家族
- 以此类推,还可能有多行连起来的,就表示这些细胞类型共享的hERV家族
- 上方的柱状图:刚才那一列点阵所定义的那个交集里,一共有多少个hERV家族
# 相关性矩阵
cor_mat <- cor(specificity_mat, method = "spearman")
df <- as.data.frame(cor_mat) %>%
tibble::rownames_to_column("celltype1") %>%
tidyr::pivot_longer(-celltype1, names_to = "celltype2", values_to = "cor")
ggplot(df, aes(x = celltype1, y = celltype2, fill = cor)) +
geom_tile(color = "grey90") +
geom_text(aes(label = sprintf("%.2f", cor)), size = 3) +
scale_fill_gradient2(low = "navy", mid = "white", high = "firebrick", midpoint = 0, limits = c(-1, 1)) +
coord_equal() +
theme_bw() +
labs(title = "Correlation of cell-type-specific hERV profiles", x = NULL, y = NULL, fill = "Spearman\nrho") +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
panel.grid = element_blank()
)
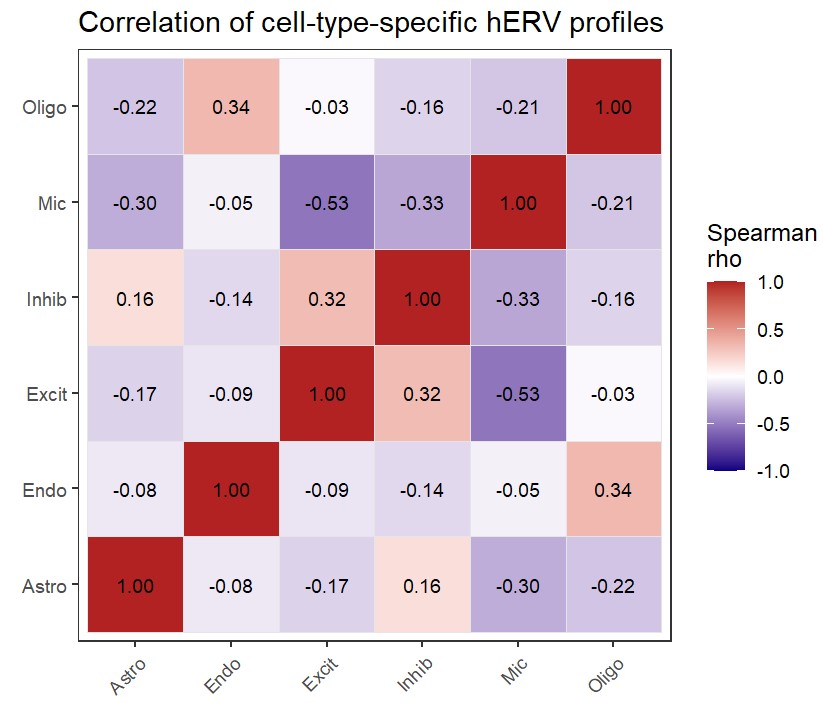
综上
- 大多数hERV家族的细胞类型特异性并不特别强,除了Mic这种有几个家族明显偏高,不过也可能是因为Mic的细胞数少导致结果不稳定
- 各细胞类型的hERV家族表达情况彼此有差异,其中Mic最独特;神经元内部(Excit/Inhib)有一定相似性,但也不是高度一致
- 单个细胞类型独有的top family占了很大比例
不过这里是把所有细胞压缩到一起看表达量,不是论文中那种每个样本算一个向量的方式
donor_col <- "orig.ident"
expr_all <- GetAssayData(seu, assay = assay_use, slot = slot_use)
meta_all <- seu@meta.data
sig_score_list <- list()
for (ct in names(top_features_by_ct)) {
feats <- top_features_by_ct[[ct]]
feats <- intersect(feats, rownames(expr_all))
ct_cells <- rownames(meta_all)[meta_all[[group_col]] == ct]
if (length(ct_cells) == 0 || length(feats) == 0) next
mat_ct <- expr_all[feats, ct_cells, drop = FALSE]
if (nrow(mat_ct) == 1) {
cell_scores <- as.numeric(mat_ct[1, ])
} else {
cell_scores <- Matrix::colMeans(mat_ct)
}
df_ct <- data.frame(
cell = ct_cells,
donor = meta_all[ct_cells, donor_col],
celltype = ct,
score = cell_scores,
stringsAsFactors = FALSE
)
donor_score_ct <- df_ct %>%
group_by(donor) %>%
summarise(score = mean(score), .groups = "drop")
colnames(donor_score_ct)[2] <- paste0(ct, "_sig")
sig_score_list[[ct]] <- donor_score_ct
}
donor_sig_df <- Reduce(function(x, y) full_join(x, y, by = "donor"), sig_score_list)
donor_sig_mat <- donor_sig_df %>%
tibble::column_to_rownames("donor") %>%
as.matrix()
cor_mat_donor <- cor(donor_sig_mat, method = "spearman", use = "pairwise.complete.obs")
df <- as.data.frame(cor_mat_donor) %>%
tibble::rownames_to_column("sig1") %>%
tidyr::pivot_longer(-sig1, names_to = "sig2", values_to = "cor")
ggplot(df, aes(x = sig1, y = sig2, fill = cor)) +
geom_tile(color = "grey90") +
geom_text(aes(label = sprintf("%.2f", cor)), size = 3) +
scale_fill_gradient2(
low = "navy", mid = "white", high = "firebrick",
midpoint = 0, limits = c(-1, 1)
) +
coord_equal() +
theme_bw() +
labs(
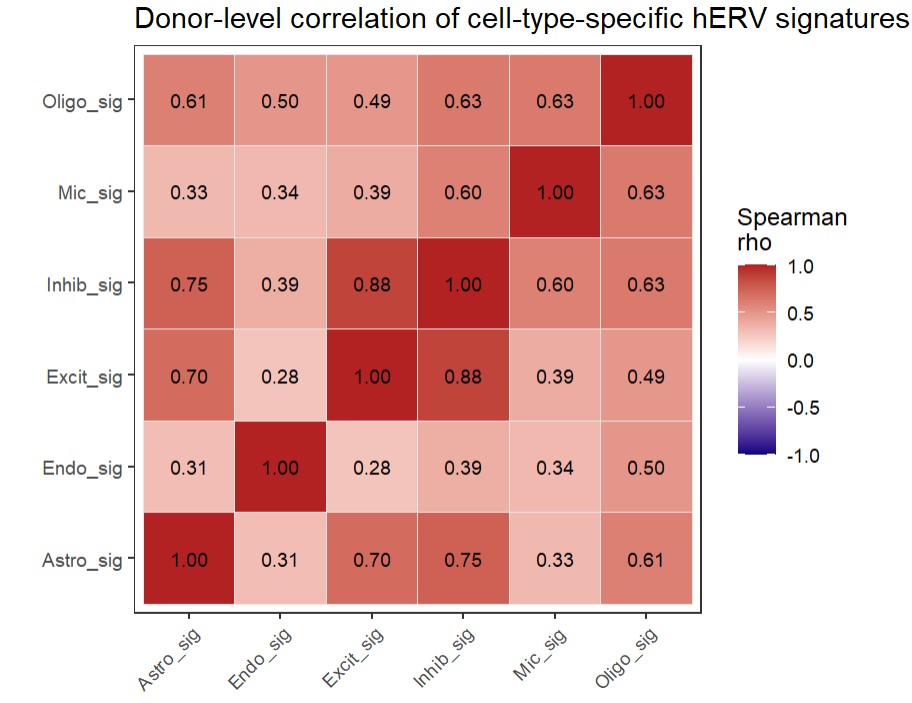
title = "Donor-level correlation of cell-type-specific hERV signatures",
x = NULL, y = NULL,
fill = "Spearman\nrho"
) +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
panel.grid = element_blank()
)
donor_sig_long <- donor_sig_df %>%
tidyr::pivot_longer(-donor, names_to = "signature", values_to = "score")
ggplot(donor_sig_long, aes(x = signature, y = score)) +
geom_violin(fill = "grey85", color = "black", trim = TRUE) +
geom_jitter(width = 0.12, size = 1.5, alpha = 0.8) +
theme_bw() +
labs(
title = "Distribution of donor-level hERV signature scores",
x = NULL, y = "Signature score"
) +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
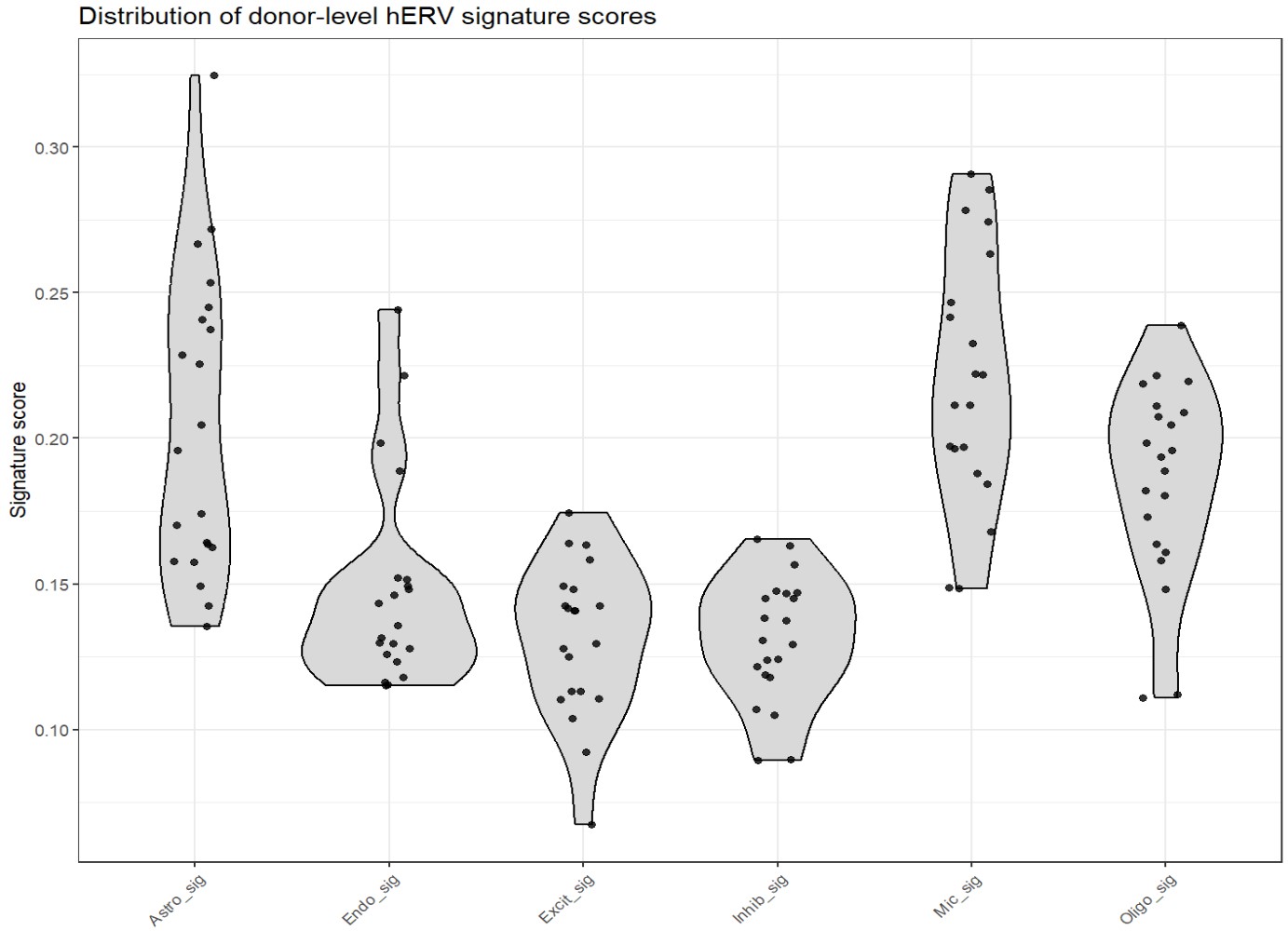
- Mic_sig、Oligo_sig、Astro_sig整体分数偏高:这几个细胞类型特异性hERV特征在donor层面更容易取得较高均值
- 其它整体偏低,说明donor间变异相对没那么大
- Astro_sig和Mic_sig的离散度比较大:donor之间差异更明显,提示这些特征更容易受个体差异影响
总的来说
- 第一张相关性图是把所有样本综合起来看,每个细胞类型的各hERV家族表达有没有相关性(不同细胞类型的hERV family构成像不像);第二张是把样本拆分看(不同donor在这些细胞类型特异性特征上是否同步变化)
- Excit和Inhib都属于神经元,所以相关性较高(某个donor如果在Excit相关hERV特征上偏高,通常在Inhib上也偏高);其它相关性都为正,说明对于大多数hERV家族,如果某样本在A高时,在B也高,感觉大概率还是测序深度的影响(即使用的是标准化后的数据),而且最主要的是我们的样本数较少,原文中的样本数都达到了k级别。也许还是用第一张相关性图说明各细胞类型的hERV组成不同 比较稳健些?